Evaluating the performance of ancient DNA genetic relatedness estimation methods using high-fidelity pedigree simulations.
Lefeuvre Maël, M Marsolier, Marie-Claude MC et al.
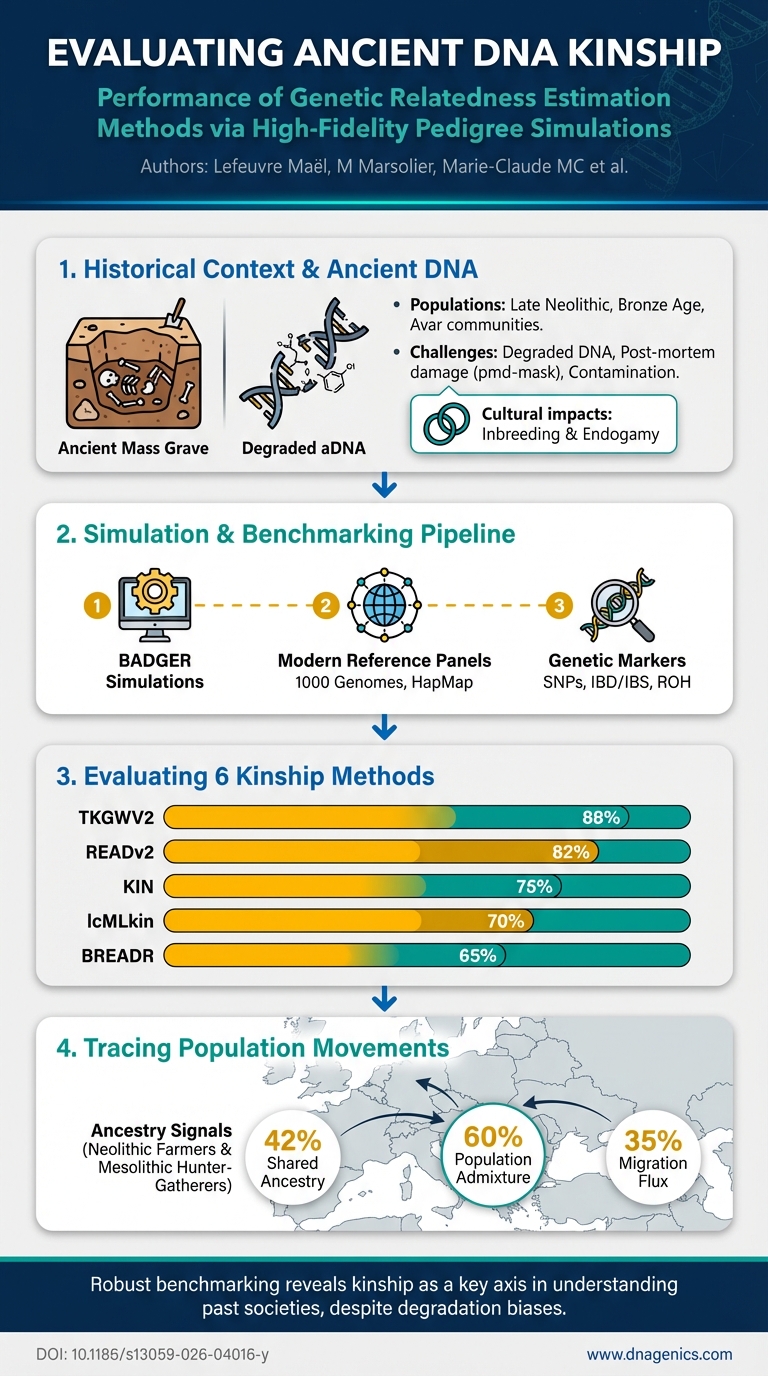
Visual Summary
A visual snapshot of this publication's key findings at a glance
Publication Details
Comprehensive information about this research publication
Abstract
Summary of the research findings
Recent advancements in paleogenetics, coupled with the emergence of dedicated statistical methods have, in recent years, streamlined the detection of close genetic ties from ancient DNA samples, leading to a substantial surge in scientific publications emphasising the reconstruction of genealogies within archaeological funerary contexts. However, while these methods all claim aptitude for addressing the inherent biases of ancient DNA, assessing their practical reliability can often be challenging, particularly in case studies involving few and/or poorly preserved samples. Furthermore, the genetic heritage and cultural practices of the population studied (e.g., inbreeding, endogamy) are factors which are often both complex to estimate and capable of impacting the accuracy of these methods.We present an in-depth comparative study of six ancient DNA genetic relatedness estimation methods to precisely delineate their respective performance and behaviour across a range of five biological parameters: sample coverage, use of post-mortem damage correction methods, human contamination, genetic diversity, and inbreeding. To this end, we develop BADGER (Benchmark Ancient DNA GEnetic Relatedness), an automated pipeline and software which first simulates pedigrees using randomly selected present-day individuals from the 1000-genomes dataset, and subsequently generates raw ancient DNA sequence data for each individual within these trees.The results of this benchmark enable us to discuss the individual strengths and limitations of these methods, propose a set of prescriptions to consider when interpreting their results and demonstrate that their reliability cannot be predicted from sample coverage alone, and may be subject to multiple sources of bias.
Listen to This Research
A two-host conversation exploring the key findings of this publication
Evaluating the performance of ancient DNA genetic relatedness estimation methods using high-fidelity pedigree simulations.
Two-Host ConversationAnalysis
Comprehensive review of ancestry and genetic findings
Important Disclaimer: This review has been performed semi-automatically and is provided for informational purposes only. While we strive for accuracy, this analysis may contain errors, omissions, or misinterpretations of the original research. DNA Genics disclaims all liability for any inaccuracies, errors, or consequences arising from the use of this information. Users should independently verify all information and consult original research publications before making any decisions based on this content. This analysis is not intended as a substitute for professional scientific review or medical advice.