A probabilistic approach to visualize the effect of missing data on PCA in ancient human genomics.
Zabel Susanne, S Breitling, Samira S et al.
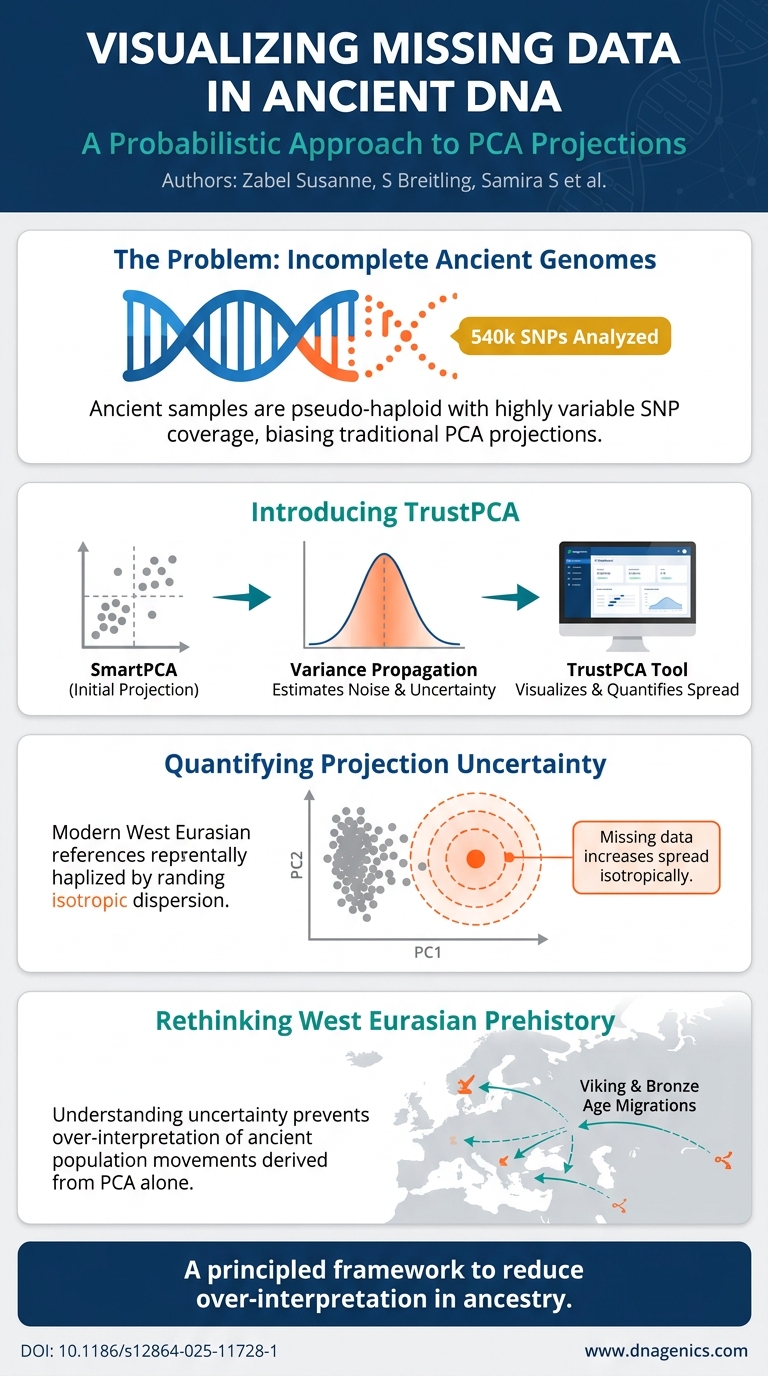
Visual Summary
A visual snapshot of this publication's key findings at a glance
Publication Details
Comprehensive information about this research publication
Abstract
Summary of the research findings
Principal Component Analysis (PCA) is widely used in population genetics to visualize genetic relationships and population structures. In ancient genomics, genotype information may in parts remain unresolved due to the low abundance and degraded quality of ancient DNA. While methods like SmartPCA allow the projection of ancient samples despite missing data, they do not quantify projection uncertainty. The reliability of PCA projections for often very sparse ancient genotype samples is not well understood. Ignoring this uncertainty may lead to overconfident conclusions about the observed genetic relationships and population structure.This study systematically investigates the impact of missing loci on PCA projections using both simulated and real ancient human genotype data. Through extensive simulations with high-coverage ancient samples, we demonstrate that increasing levels of missing data can lead to less accurate SmartPCA projections, highlighting the importance of considering uncertainty when interpreting PCA results from ancient samples. To address this, we developed a probabilistic framework to quantify the uncertainty in PCA projections due to missing data. By applying our methodology to modern and ancient West Eurasian genotype samples from the Allen Ancient DNA Resource database, we could show a high concordance between our predicted projection and empirically derived distributions. Applying this framework to real-world data, we demonstrate its utility in predicting and visualizing embedding uncertainties for ancient samples of varying SNP coverages.Our results emphasize the importance of accounting for projection uncertainty in ancient population studies. We therefore make our probabilistic model available through TrustPCA, a user-friendly web tool that provides researchers with uncertainty estimates alongside PCA projections, facilitating data exploration in ancient human genomic studies and enhancing transparency in data quality reporting.
Listen to This Research
A two-host conversation exploring the key findings of this publication
A probabilistic approach to visualize the effect of missing data on PCA in ancient human genomics.
Two-Host ConversationAnalysis
Comprehensive review of ancestry and genetic findings
Important Disclaimer: This review has been performed semi-automatically and is provided for informational purposes only. While we strive for accuracy, this analysis may contain errors, omissions, or misinterpretations of the original research. DNA Genics disclaims all liability for any inaccuracies, errors, or consequences arising from the use of this information. Users should independently verify all information and consult original research publications before making any decisions based on this content. This analysis is not intended as a substitute for professional scientific review or medical advice.