A scalable pipeline for local ancestry inference using tens of thousands of reference haplotypes
Durand, E. Y., Do, C. B., Wilton, P. R. et al.
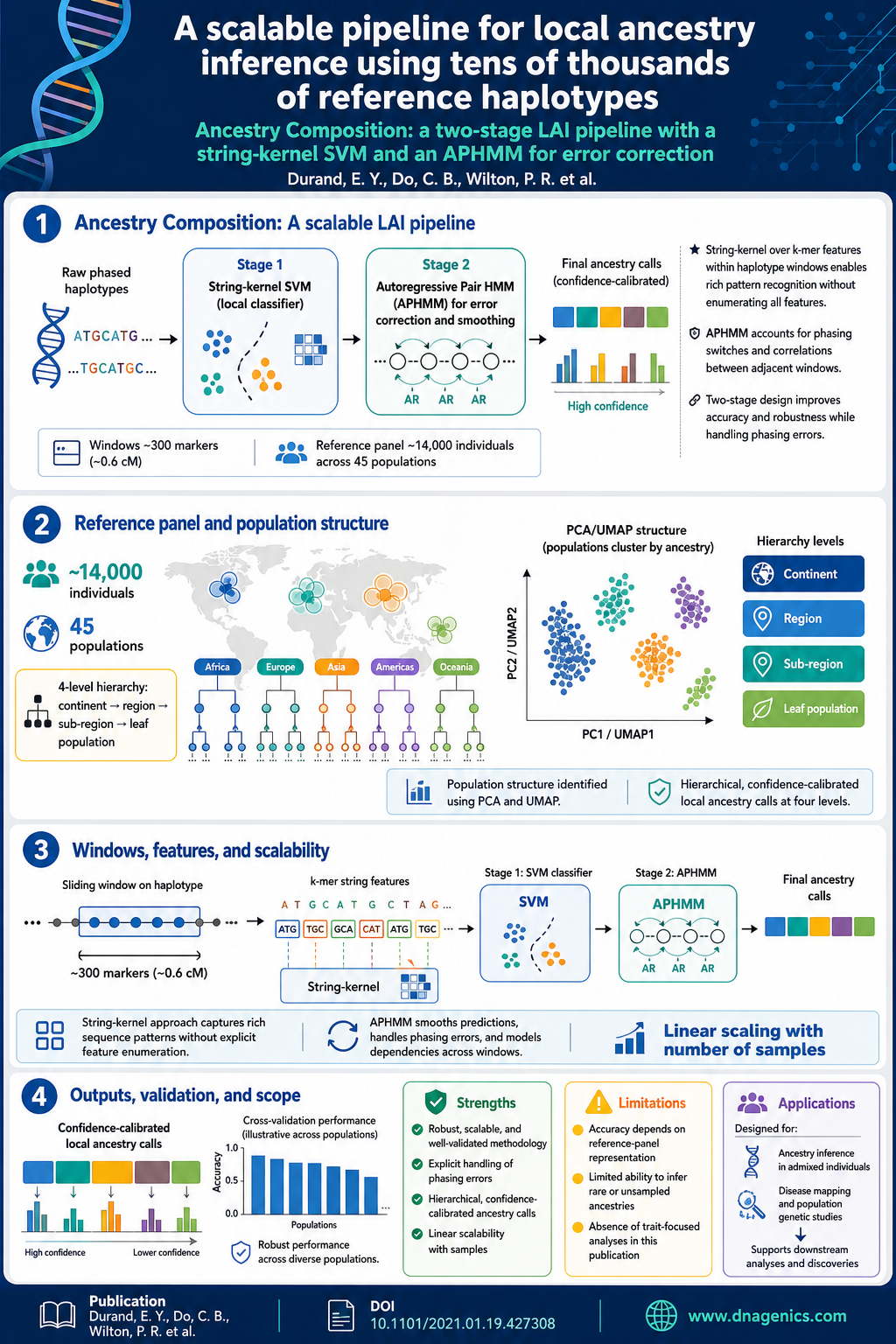
Visual Summary
A visual snapshot of this publication's key findings at a glance
Publication Details
Comprehensive information about this research publication
Abstract
Summary of the research findings
Ancestry deconvolution is the task of identifying the ancestral origins of chromosomal segments of admixed individuals. It has important applications, from mapping disease genes to identifying loci potentially under natural selection. However, most existing methods are limited to a small number of ancestral populations and are unsuitable for large-scale applications. In this article, we describe Ancestry Composition, a modular pipeline for accurate and efficient ancestry deconvolution. In the first stage, a string-kernel support-vector-machines classifier assigns provisional ancestry labels to short statistically phased genomic segments. In the second stage, an autoregressive pair hidden Markov model corrects phasing errors, smooths local ancestry estimates, and computes confidence scores. Using publicly available datasets and more than 12,000 individuals from the customer database of the personal genetics company, 23andMe, Inc., we have constructed a reference panel containing more than 14,000 unrelated individuals of unadmixed ancestry. We used principal components analysis (PCA) and uniform manifold approximation and projection (UMAP) to identify genetic clusters and define 45 distinct reference populations upon which to train our method. In cross-validation experiments, Ancestry Composition achieves high precision and recall.
Listen to This Research
A two-host conversation exploring the key findings of this publication
A scalable pipeline for local ancestry inference using tens of thousands of reference haplotypes
Two-Host ConversationAnalysis
Comprehensive review of ancestry and genetic findings
Important Disclaimer: This review has been performed semi-automatically and is provided for informational purposes only. While we strive for accuracy, this analysis may contain errors, omissions, or misinterpretations of the original research. DNA Genics disclaims all liability for any inaccuracies, errors, or consequences arising from the use of this information. Users should independently verify all information and consult original research publications before making any decisions based on this content. This analysis is not intended as a substitute for professional scientific review or medical advice.