Introduction
In the medieval heart of Berlin, the cemetery at St. Peter’s Churchyard in Cölln sits across the Spree from what would become Germany’s capital. Researchers opened a window into the lives of people who lived there between roughly 1150 and 1717 by analyzing bones and teeth with both genetic and isotopic tools. This combination — ancient DNA markers and dental enamel isotopes — lets us peek at where individuals grew up, who their ancestors were, and how much mobility shaped the community.
Why this matters goes beyond curiosity about history. By linking genetic signals with geographic provenance, the study illuminates long-term population dynamics in one of Europe’s oldest urban nuclei. The findings contribute to debates about how medieval populations formed, moved, and integrated in a region that experienced Slavic influences, Germanic settlement during the Ostsiedlung, and sustained local continuity. The St. Peter’s sample provides a case study for how multiple lines of evidence converge to outline ancestry, migration, and social structure in a key historical landscape.
Key Discoveries
- Multi-proxy dataset: The study combines genetic data (autosomal STRs, Y-STRs, mtDNA) and isotopic analyses (strontium and oxygen in dental enamel) from a medieval cemetery in Berlin–Cölln.
- Y-haplogroup mix: Males show a distribution dominated by R1b and R1a, with I1 and I2a also present, patterns compatible with Central/Northern European male lineages.
- mtDNA continuity: Maternal lineages cluster around European haplogroups H, U, T, J, and K.
- Local mobility signal: Isotope data indicate most individuals originated in the Berlin-Brandenburg region, with only a minority showing signals of migration.
- Methodological caveats: STR-based analyses are informative for kinship and recent ancestry but have limited power for deep ancestry; deeper resolution benefits from genome-wide SNP data and dedicated Y/mtDNA SNP sequencing.
- Interpretation caution: Assigning modern ethnic labels (e.g., Slavic vs Germanic) to haplogroups is risky and should be grounded in broad historical context and temporal sampling.
What This Means for Your DNA
For people exploring their own ancestry, this study reinforces a few practical takeaways. First, ancestry signals in medieval contexts are often local: many individuals in a sustained urban population show genetic ties to the surrounding region across generations. Second, different genetic markers tell different stories. Autosomal STRs capture kinship and nearby relatedness, while Y-chromosome and mtDNA markers reveal paternal and maternal lineage patterns, respectively. Third, isotopic data — such as strontium and oxygen in enamel — provide geographic anchors for where a person likely grew up, enriching genetic inferences with a geography of upbringing.
For consumer DNA analysis, the lesson is not that haplogroups alone define ancestry, but that combining genetic signals with geographic context and, when possible, historical data yields a richer narrative. Modern DNA tests can tell you about your broader ancestral regions, but deeper population history often requires broader genomic data and careful interpretation within archaeological contexts. This study illustrates why multi-faceted approaches produce the most robust picture of medieval populations and how those lessons translate to contemporary DNA analyses.
Historical and Archaeological Context
The St. Peter’s cemetery in Cölln operated from about 1150 to 1717, a period that tracks the emergence of Berlin as a market and urban center and the broader arc of Germanic settlement in eastern Europe. The genetic and isotopic results align with known historical movements: a substantial local continuity in the Berlin-Brandenburg region, paired with signals of modest migration that fit the city’s growth and trade links at various points in the medieval and early modern eras.
From an archaeological perspective, the combination of autosomal, Y-chromosome, and mtDNA markers with enamel isotopes provides a layered view of population dynamics. The prevalence of Central/Northern European paternal lineages and European maternal lineages suggests a complex but regionally grounded demographic history, reflecting long-standing local residence punctuated by small-scale movements. The timeline and geography underscore how urban centers like Berlin–Cölln functioned as hubs where local populations persisted and occasional migrants contributed to the community fabric.
The Science Behind the Study
This study employs a robust, multi-proxy methodology suitable for readers who want to understand how ancient population histories are reconstructed. The researchers analyzed 96 individuals for 16 autosomal STRs plus the amelogenin marker to determine sex. They identified 54 unrelated males and typed them for 27 Y-STRs, assigning them to 12 Y-haplogroups. Geographic ancestry predicted by AMOVA showed strong similarity to present-day Germany, with notable differences when compared to neighboring western and eastern European populations. Mitochondrial DNA typing was performed on all 96 individuals using coding-region SNPs, revealing a typical European distribution of haplogroups H, U, T, J, and K. A subset of 48 individuals underwent full mtDNA control-region sequencing, which indicated only trace amounts of Slavic-associated lineages. In parallel, strontium and oxygen isotope analyses were performed on 66 of the earliest graves, indicating most individuals originated from the Berlin–Brandenburg area with limited migration signals.
- The sample sizes (96 individuals for STRs and mtDNA; 54 Y-chromosome–typed males; 66 isotope analyses) provide a solid basis for regional inferences, though the authors caution about limitations.
- The methodology relies on targeted marker sets (STRs and SNPs for mtDNA) rather than genome-wide data, which means conclusions about deep ancestry and fine-scale admixture are inherently limited compared to modern ancient genome-wide studies.
In Simple Terms: STRs are short DNA repeats that are great for identifying who is related to whom in the near past (like siblings or cousins) but not as precise for mapping very old ancestry. The Y-chromosome SNPs and mtDNA variants track paternal and maternal lines, respectively, while isotope data tell you where a person likely grew up. Together, they form a fuller picture than any single method alone, but genome-wide SNP data would offer an even sharper view of ancient population history.
Infographic Section
To visually summarize the study, see the infographic below. It highlights the multi-proxy approach (genetics and isotopes), the distribution of Y-haplogroups and mtDNA haplogroups, and the local origin signal from isotope data.
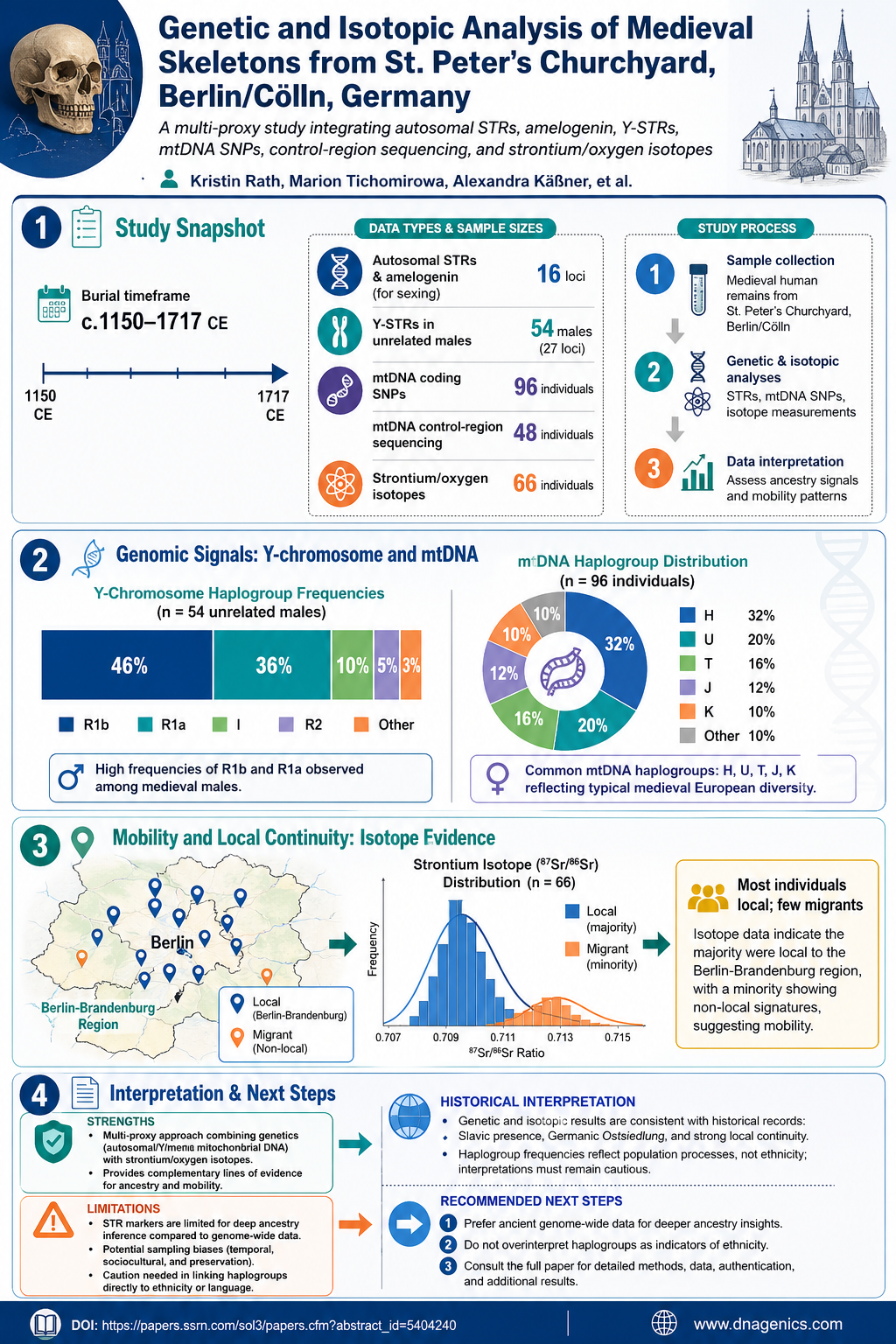

What the infographic shows:
- The proportion of different Y-haplogroups among the 54 male individuals, with R1b and R1a as dominant lineages.
- The mtDNA haplogroup distribution across the population, emphasizing European lineages H, U, T, J, and K.
- Isotopic evidence indicating most individuals originated in the local region (Berlin-Brandenburg), with a subset suggesting migration.
- A reminder of the study’s multi-proxy design and its limitations.
Why It Matters
This research contributes to our understanding of medieval population dynamics in a pivotal European urban center. It demonstrates how combining genetic data with isotopic signatures can distinguish local continuity from mobility, a key question for population genetics and historical archaeology. The findings reinforce that while some early Germanic and Slavic influences may have left traces, much of the cemetery’s population appears locally rooted, aligning with known regional history. Looking ahead, integrating ancient genome-wide data with broader temporal and geographic sampling will sharpen our capacity to resolve population structure, migration waves, and cultural interactions in medieval Europe.
Future work could expand genome-wide analyses to create higher-resolution population histories, compare additional cemeteries across the region, and refine methods to interpret haplogroup signals within historical contexts. This study sets a strong precedent for multi-proxy approaches in archaeology-driven genetics and underscores the importance of cautious interpretation when linking genetic markers to ethnicity or nationality.