Introduction
Ancient genomes carry invisible threads linking people across time. Long DNA segments shared between individuals, known as identity-by-descent (IBD), reveal recent genealogical connections even when data are sparse. This makes it possible to read kinship signals that archaeology alone cannot fully resolve.
This study introduces ancIBD, a specialized method for detecting IBD segments in ancient DNA by combining a five-state hidden Markov model with imputed genotype probabilities. By leveraging low-coverage data and a reference haplotype panel, ancIBD can recover meaningful kinship signals where traditional analyses struggle, providing a bridge between personal genealogies and broad cultural migrations.
Key Discoveries
- ancIBD detects IBD segments longer than ~8 cM in low-coverage ancient DNA, enabling robust kinship inferences.
- In a dataset of 4,248 ancient Eurasian individuals, relatives up to the sixth degree were identified.
- Corded Ware–Yamnaya co-ancestry is evident within a few generations, highlighting recent co-migration or intermarriage links.
- Afanasievo individuals buried ~1,410 km apart show long IBD segments, indicating recent cross-regional kinship or mobility.
- WGS data outperform 1240k data at the same coverage, and imputation at 1240k sites provides substantial off-target data advantages; ancIBD balances precision and recall better than other methods under low coverage.
What This Means for Your DNA
Ancient DNA studies like this illuminate how kinship networks formed, migrated, and changed over time. For researchers and enthusiasts, the work demonstrates that meaningful genealogical inferences can be drawn even from sparse data, provided robust imputation and probabilistic modeling are used. While the immediate application is to ancient genomes, the underlying principle—detecting long shared blocks to infer recent ancestry—resonates with modern DNA analyses as well.
For those exploring ancestry more broadly, the study underscores two practical takeaways: (1) data depth matters. Whole-genome sequencing (WGS) offers clearer IBD signals than targeted SNP capture when depth is comparable. (2) Imputation can unlock information from low-coverage data, but it relies on reference panels; researchers should interpret results within the context of available panels and archaeological background. In the future, integrating ancIBD-style approaches with consumer-genetics workflows could enrich interpretations of ancient kinship alongside modern lineages.
Historical and Archaeological Context
The ancIBD findings dovetail with long-standing models of Steppe-driven migrations and Bronze Age admixture across Europe and Eurasia. By identifying close and distant relatives among Corded Ware, Yamnaya, Afanasievo, and Globular Amphora contexts, the work provides direct genealogical support for proposed demographic links between Pontic-Caspian Steppe herders and European populations during the late third millennium BCE. The detected long-range IBD sharing between Corded Ware and Yamnaya groups suggests substantial co-ancestry within a few generations, reinforcing narratives of rapid cultural and genetic exchange across vast regions.
Across space, Afanasievo individuals buried thousands of kilometers apart show long IBD segments, signaling mobility and kinship that transcended local groups. Together, these patterns illuminate where kin networks overlapped with migrations, helping historians and archaeologists map how kinship ties shaped cultural contact, trade, and settlement patterns during a dynamic era of population turnover.
The Science Behind the Study
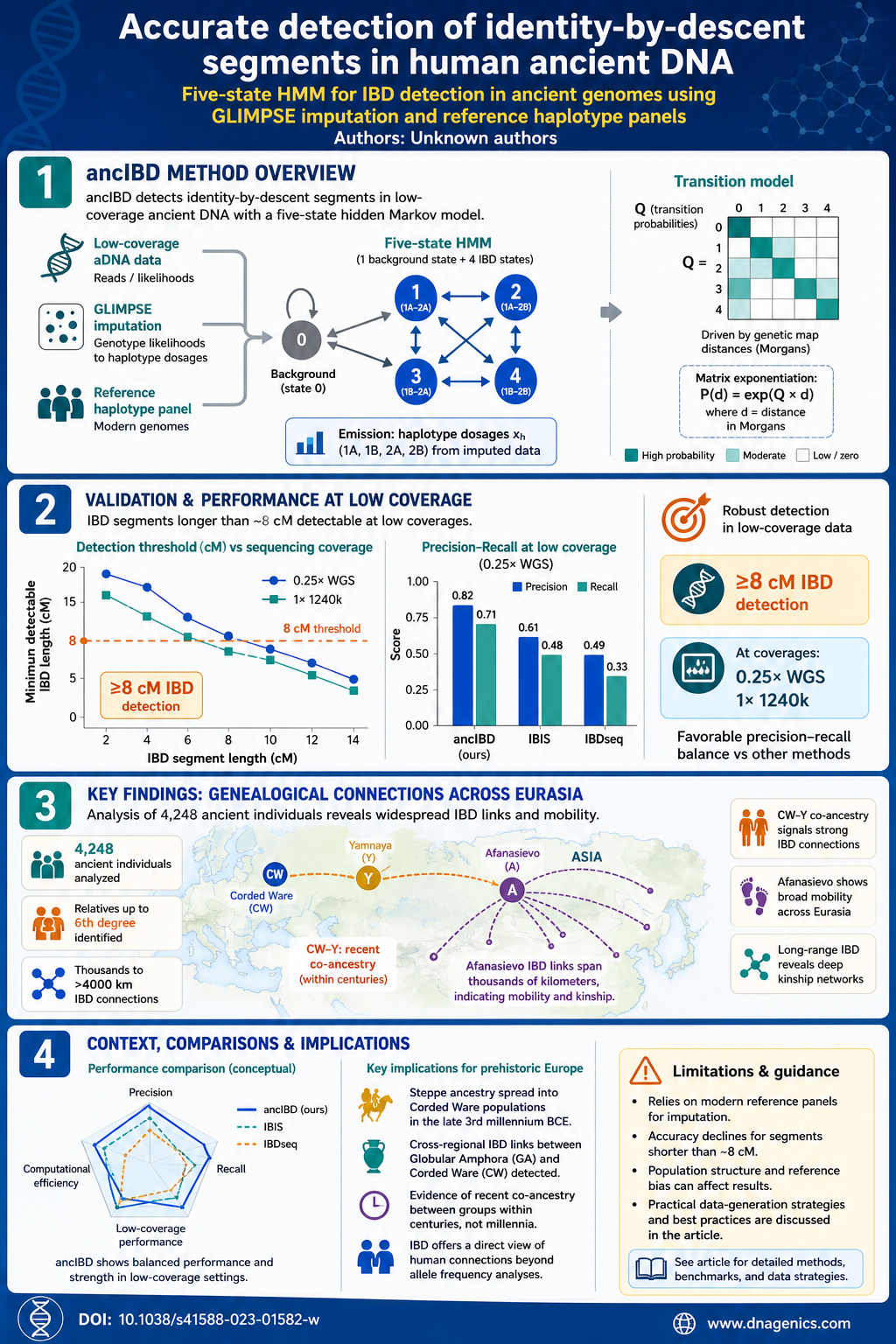
ancIBD uses a five-state hidden Markov model with one background state and four IBD-sharing states (representing the four possible haplotype-pairings). Emission probabilities are computed from imputed haplotype dosages, while transition probabilities are driven by genetic map distances (Morgans) through a parameter matrix Q and matrix exponentiation. The genotype data are first imputed with GLIMPSE to generate probabilistic haplotypes suitable for detecting IBD in low-coverage data. The approach is validated on both whole-genome sequencing (WGS; ~0.25×) and 1240k SNP capture data at comparable depths.
The study demonstrates that ancIBD robustly identifies IBD segments longer than ~8 cM under low-coverage conditions and that WGS data offer better performance than 1240k data at the same coverage, though 1240k imputation still provides meaningful off-target information. When compared to existing methods like IBIS and IBDseq, ancIBD achieves a favorable balance of precision and recall in low-coverage ancient DNA scenarios. The authors also discuss limitations, such as dependence on modern reference panels for imputation, and provide practical guidance for data-generation strategies.
In Simple Terms: ancIBD is like a detective’s clock for old genomes. It uses a probabilistic model and a reference map of shared DNA to find long blocks that indicate recent kinship, even when the data are noisy or sparse.
[Infographic Section - Infographic: ancIBD in ancient genomes – kinship and migration insights]
The accompanying infographic summarizes how ancIBD works, the data depth where it performs best, and the major findings from the Eurasian dataset, including Corded Ware–Yamnaya co-ancestry and Afanasievo mobility signals. It visualizes the 8 cM detection threshold, the scale of the sample (4,248 ancient genomes), and the key population-level connections uncovered by the method.

Why It Matters
This work advances our ability to read the genealogical clock in ancient genomes, converting long-standing questions about kinship and migration into testable signals. By enabling reliable IBD detection in low-coverage aDNA, ancIBD broadens the scope of what archaeologists and population geneticists can infer about population structure, mobility, and cultural interactions in prehistory. Future directions include refining imputation with ancient reference panels, expanding the coverage of ancient genomes, and integrating IBD-based kinship signals with archaeological and isotopic data to build richer demographic reconstructions.
References
Accurate detection of identity-by-descent segments in human ancient DNA
- DOI: https://doi.org/10.1038/s41588-023-01582-w