Introduction
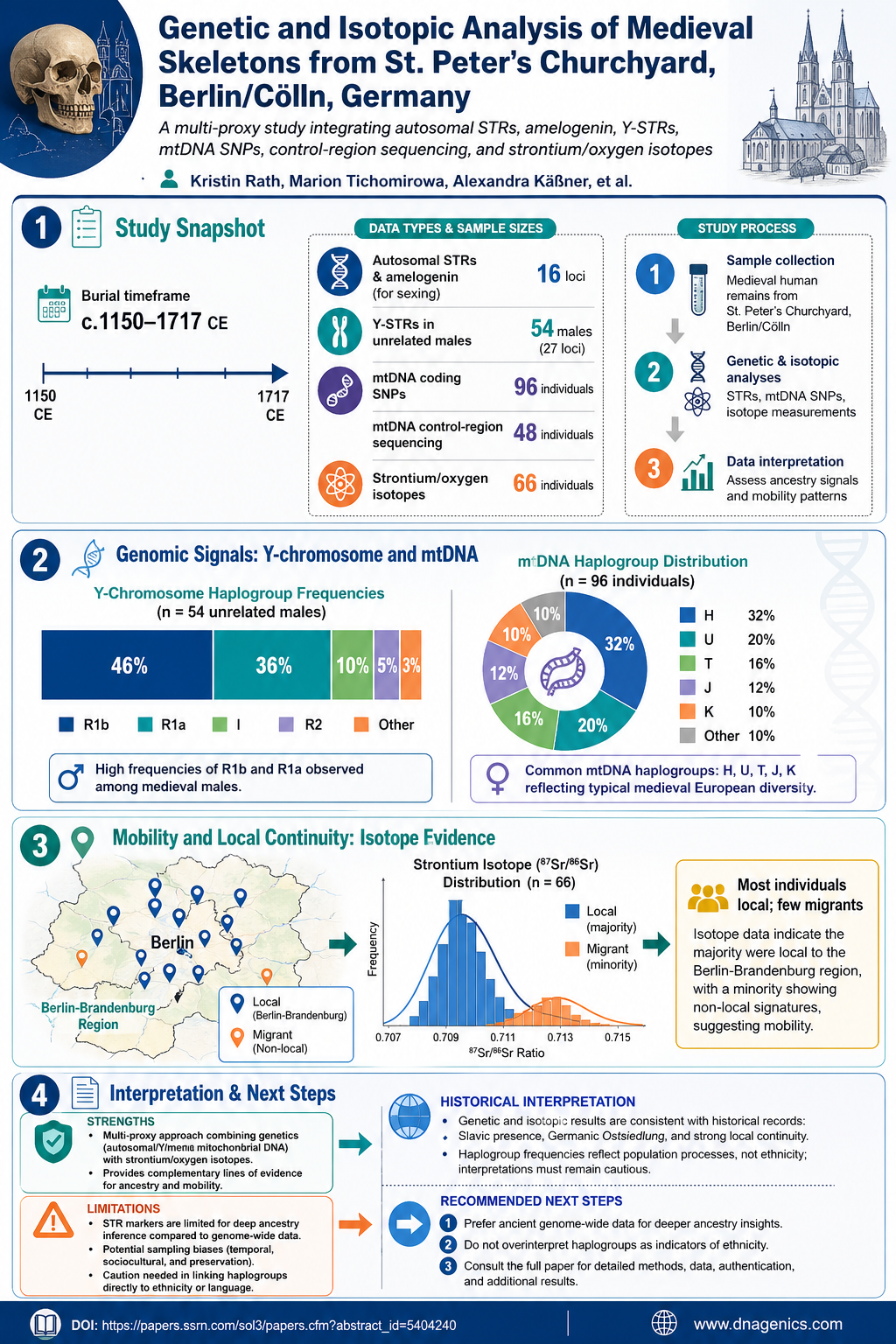
Mitochondrial DNA (mtDNA) holds a compact, maternally inherited record of our ancestry. Yet analyzing mtDNA has long faced a stubborn obstacle: NUMTs, fragments of mitochondrial origin embedded in the nuclear genome, can confound sequencing and interpretation. This study tackles that challenge with a novel nanopore-based third-generation sequencing (TGS) approach that uses a single primer pair to amplify the full-length mtDNA, reducing NUMT artifacts and boosting data clarity.
By sequencing full-length mtDNA across 106 samples from eight family pedigrees—encompassing complex structures like half-siblings and multi-generational households—the researchers achieved 100% genome coverage and an impressive mean depth of around 1896×. Sequencing on the QITAN TECH QNome-3841hex platform produced high-quality reads with a mean mapping rate of 99.96%, enabling robust maternal lineage validation and comprehensive mtDNA characterization. The resulting dataset promises meaningful advances in disease research, evolutionary studies, ancestry tracing, and forensic identification, while also guiding future improvements in mtDNA sequencing technology.
This post introduces the study’s breakthroughs, what they mean for your DNA analysis, and how they fit into the broader scientific and historical context of maternal inheritance and human migration.
Key Discoveries
- Full-length mtDNA sequencing of 106 individuals across eight pedigrees was achieved with 100% genome coverage and mean depth ≈1896×, enabling high-confidence haplotype reconstruction.
- Maternal lineage concordance is strong: 73 of 74 maternally related samples showed concordant haplogroups (≈98.6%), supporting reliable maternal inheritance signals.
- NUMT interference minimized by a single-amplicon long-range PCR design and length filtering; 36 samples showed some mtDNA–nuclear read overlap (78 events) but overall mtDNA purity remains high.
- Haplogroup classification and phylogenetic analyses rely on HaploGrep 3 and Phylotree Build 17 with rCRS normalization, supporting robust ancestry inferences within the dataset's scope.
- Open data and reproducible workflows (SRA, GenBank, EVA, figshare, GitHub) enable reuse for forensic mtDNA genotyping, maternal lineage tracing, and population genetics, with appropriate caution about heteroplasmy and NUMTs.
Ancestry Insights
- Maternal lineage signals: Strong alignment of mtDNA haplotypes within maternal lines across eight families.
- Haplogroup confidence: High-confidence calls using HaploGrep 3 and Phylotree Build 17 bolster ancestry inferences.
- Data transparency: Open-data approach supports independent validation and method development.
- Forensic relevance: A well-characterized, full-length mtDNA resource enhances maternal lineage tracing and identification in forensics.
Historical and Archaeological Context
The study’s full-length mtDNA data deepen our understanding of maternal inheritance across generations and align with established migration narratives. By resolving mtDNA haplogroups with a standardized reference framework (rCRS) and robust phylogenetic trees (Phylotree Build 17), the dataset anchors maternal lineages to broader population histories. While the eight pedigrees represent modern family structures, their mtDNA haplotypes can reflect ancient migrations that shaped contemporary distributions.
Phylogenetic clustering consistent with known maternal relationships confirms expected maternal continuities within families and mirrors how mtDNA has been used to trace migrations in archaeology and anthropology. The combination of high-coverage sequencing and transparent data practices also facilitates cross-study comparisons, enabling researchers to place these intergenerational mtDNA patterns within global timelines of human movement.
The Science Behind the Study
The authors introduce a targeted, single-primer long-range PCR strategy to amplify the entire mitochondrial genome in one amplicon, followed by long-read nanopore sequencing on the QITAN TECH QNome-3841hex platform. This design minimizes NUMT interference that commonly plagues short-fragment approaches and permits unbroken mtDNA reads, resulting in near-complete genome coverage (100%) and a mean depth of approximately 1896× across 106 samples. The study reports a robust 99.96% mapping rate, underscoring the data’s reliability for downstream haplogroup assignment and phylogenetic analysis.
For haplogroup calls, the researchers employed HaploGrep 3 and Phylotree Build 17 with rCRS normalization, ensuring that classifications align with contemporary haplogroup nomenclature and phylogenetic structure. They conducted careful NUMT filtering and length-based read filtering to further reduce nuclear contamination, acknowledging limitations such as mega-NUMTs and the intrinsic error profiles of nanopore sequencing. Importantly, the dataset is openly accessible (SRA, GenBank, EVA, figshare) and accompanied by reproducible workflows (GitHub), enabling reuse in forensic mtDNA work, maternal lineage tracing, and population genetics.
In Simple Terms: The study uses one long DNA primer to copy the entire mitochondrial genome in a single piece, then reads it with a long-read technology. This helps avoid confusing pieces of nuclear DNA that resemble mitochondria, making it easier to read the true maternal DNA and trace family history. The data are shared openly so other scientists can verify results or reuse the methods.
[Infographic Section - Infographic Available]
The infographic summarizes the study design, sequencing workflow, and data availability, illustrating how a single-primer, long-range mtDNA amplification feeds into nanopore sequencing, with open-data resources wrapping up the workflow for reproducibility.
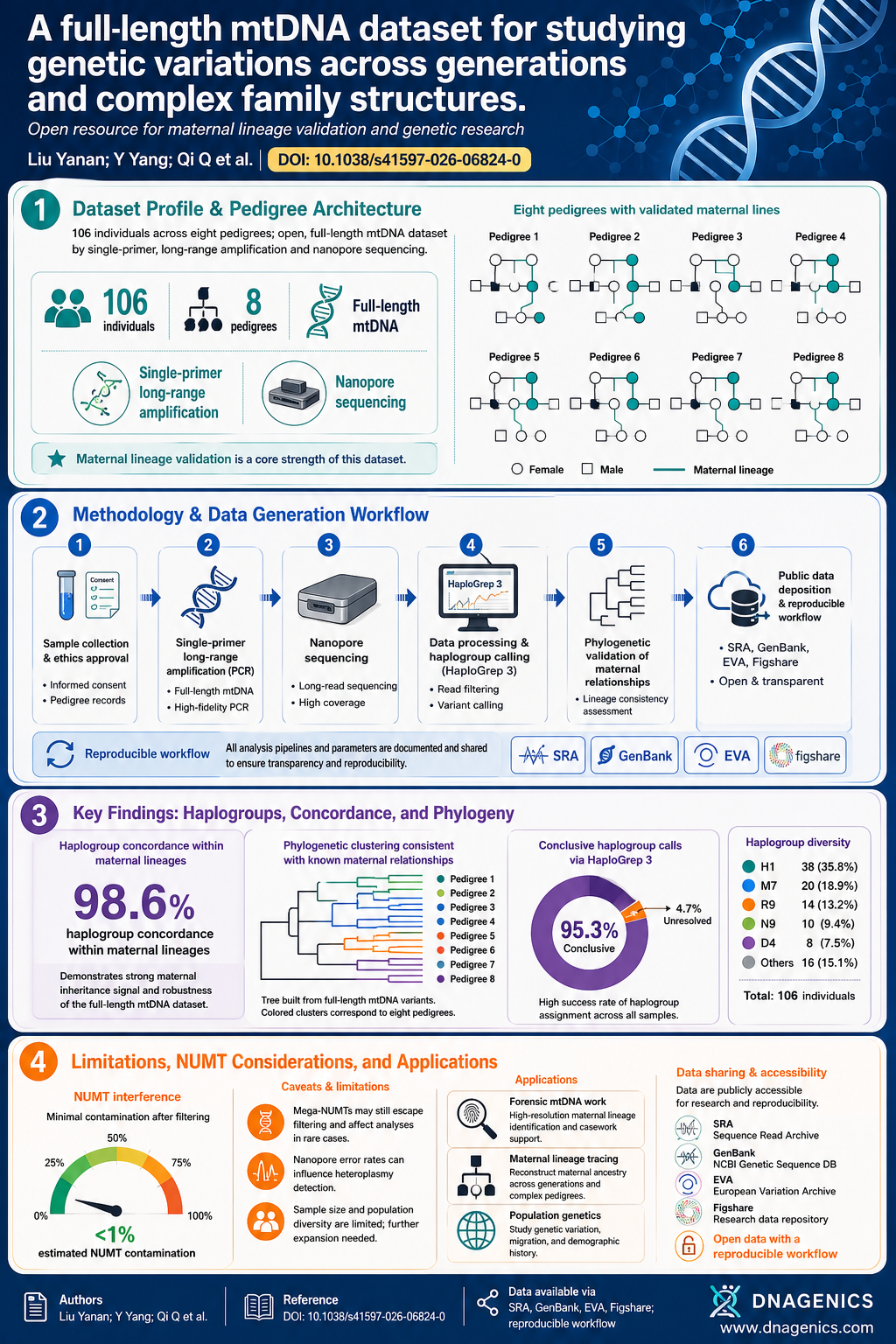

Caption: The infographic visualizes the single-primer, long-range mtDNA amplification, nanopore sequencing workflow, and open data resources that support reproducible ancestry research.
Why It Matters
This work advances mtDNA sequencing by delivering a high-coverage, full-length mtDNA resource with robust maternal lineage validation and transparent, reusable workflows. The open-data framework lowers barriers to replication, forensic validation, and cross-study comparisons in population genetics and ancestry research. Future directions include expanding haplotype diversity, addressing mega-NUMTs more comprehensively, and integrating these full-length mtDNA datasets with nuclear genome data to enrich intergenerational studies and forensic practice.
References
DOI: 10.1038/s41597-026-06824-0