Introduction
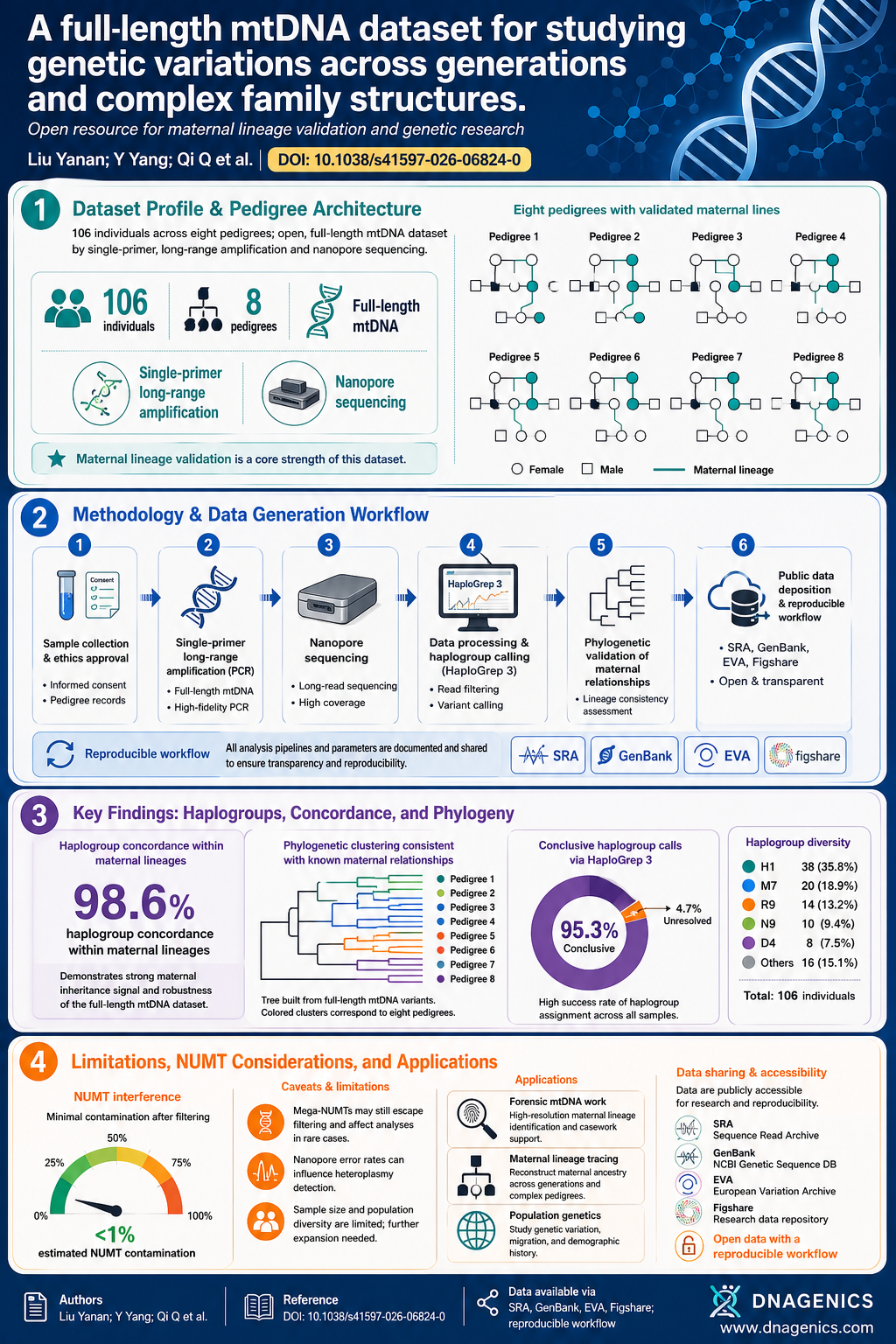
Ancient genomes are fragile clues to our past, often preserved only as fragments. Yet those fragments can reveal a family story—who among the dead was related, who shared a cell lineage, and how kinship shaped burial practices. Enter KIN, a method designed to infer relatedness from low-coverage ancient DNA by looking at shared identical-by-descent (IBD) segments. This approach makes kinship analyses feasible even when data are sparse, noisy, or contaminated.
This research matters because reconstructing kin networks in ancient populations helps us understand social structure, lineage, and population dynamics, enriching interpretations of past cultures and migrations. The study also provides practical tools to support downstream genetic analyses that depend on knowing who is related. The work sits within a wider era of archaeogenomics that includes Bronze Age Europe, steppe migrations, and Neanderthal genetics, offering a concrete method to illuminate family ties in the archaeological record.
Note: this is a preprint. Results depend on data quality, reference allele frequencies, and ancient DNA damage patterns, and the study does not report trait predictions. The authors also share that AI analysis is available to summarize findings, highlighting the method’s accessibility for diverse readers and researchers.
Key Discoveries
-
- IBD segments shared between two ancient individuals enable kinship inference, with reliable classification up to 3rd-degree relatives at a minimum 0.05x sequence coverage.
-
- Siblings vs parent-child relationships can be distinguished, adding nuance to social interpretation.
-
- Contamination and inbreeding models improve classification accuracy, addressing common biases in ancient DNA data.
-
- The method clarifies kin networks within populations, enriching understanding of ancient social structure.
What This Means for Your DNA
For researchers and enthusiasts tracing ancestry through ancient remains, KIN offers a practical route to reconstruct family ties even when DNA data are scarce. The ability to infer relationships up to the third degree from low-coverage genomes means you can map broader kin networks in a population and place individuals within their family context. While KIN is tailored for ancient DNA, the underlying emphasis on IBD-based kinship and careful handling of contamination echoes principles used in modern forensic and genealogical studies, illustrating how ancient and modern genetics converge.
Practically, when working with ancient samples, expect to report not only who is related but the confidence in those relationships, given the coverage and data quality. KIN’s framework also demonstrates the importance of accounting for contamination and inbreeding, which can otherwise blur kinship signals. For those using DNA data to explore ancestry, the study underscores the value of integrating kinship inference with population context to interpret past lifeways more accurately.
AI analysis is available for this publication, offering rapid summaries and interpretations of the kinship inferences, which can be helpful for researchers and educators communicating results to broader audiences.
Historical and Archaeological Context
Kinship information from ancient DNA enriches interpretations of burial customs, lineage-based status, and group dynamics. The ability to detect up to third-degree relatives within ancient populations provides a more nuanced view of social organization and how families organized themselves within communities. This aligns with broader archaeological inquiries into how kinship shaped ritual practices, inheritance, and settlement patterns, particularly in regions affected by Bronze Age transitions and steppe migrations.
By situating kinship signals within known migration routes and cultural exchanges, researchers can correlate genetic connections with archaeological artifacts, settlement layouts, and grave goods. The method advances our capacity to link genetic data with material culture, helping to timeline-harmonize when and where kin-based social structures emerged and how they evolved over time.
The Science Behind the Study
Kinship inference with KIN rests on analyzing identical-by-descent (IBD) segments between pairs of ancient genomes. IBD segments are stretches of DNA shared because of a recent common ancestor. In low-coverage data, direct genotype calls are unreliable, so KIN employs probabilistic approaches to detect IBD evidence despite missing data and DNA damage typical of ancient samples. The framework models how contamination and inbreeding can distort signals, allowing more accurate classification of relationships up to the 3rd degree and the ability to tell apart siblings from parent-child pairs.
The study leverages both simulated datasets and empirical ancient DNA data to validate performance across different coverage levels, with a core emphasis on the ≥ 0.05x coverage threshold. By incorporating contamination-adjustment and inbreeding-detection components, KIN improves both sensitivity and specificity in identifying kinship, providing a robust tool for parsing kin networks in ancient populations.
In Simple Terms: Kinship here means family ties. The method looks for DNA chunks that two individuals share because they inherited them from the same ancestor. Even with only tiny bits of data (0.05x coverage), KIN can tell who was closely related and whether two people were siblings or parent-child, while also accounting for sample contamination and inbreeding that could skew results.
[Infographic Section - ONLY if infographic is available]
The infographic visually summarizes how KIN detects IBD segments, the coverage threshold, and how relationships are categorized. It illustrates the workflow from data quality, through IBD detection, to kinship classification, and highlights how contamination and inbreeding are modeled to improve accuracy.
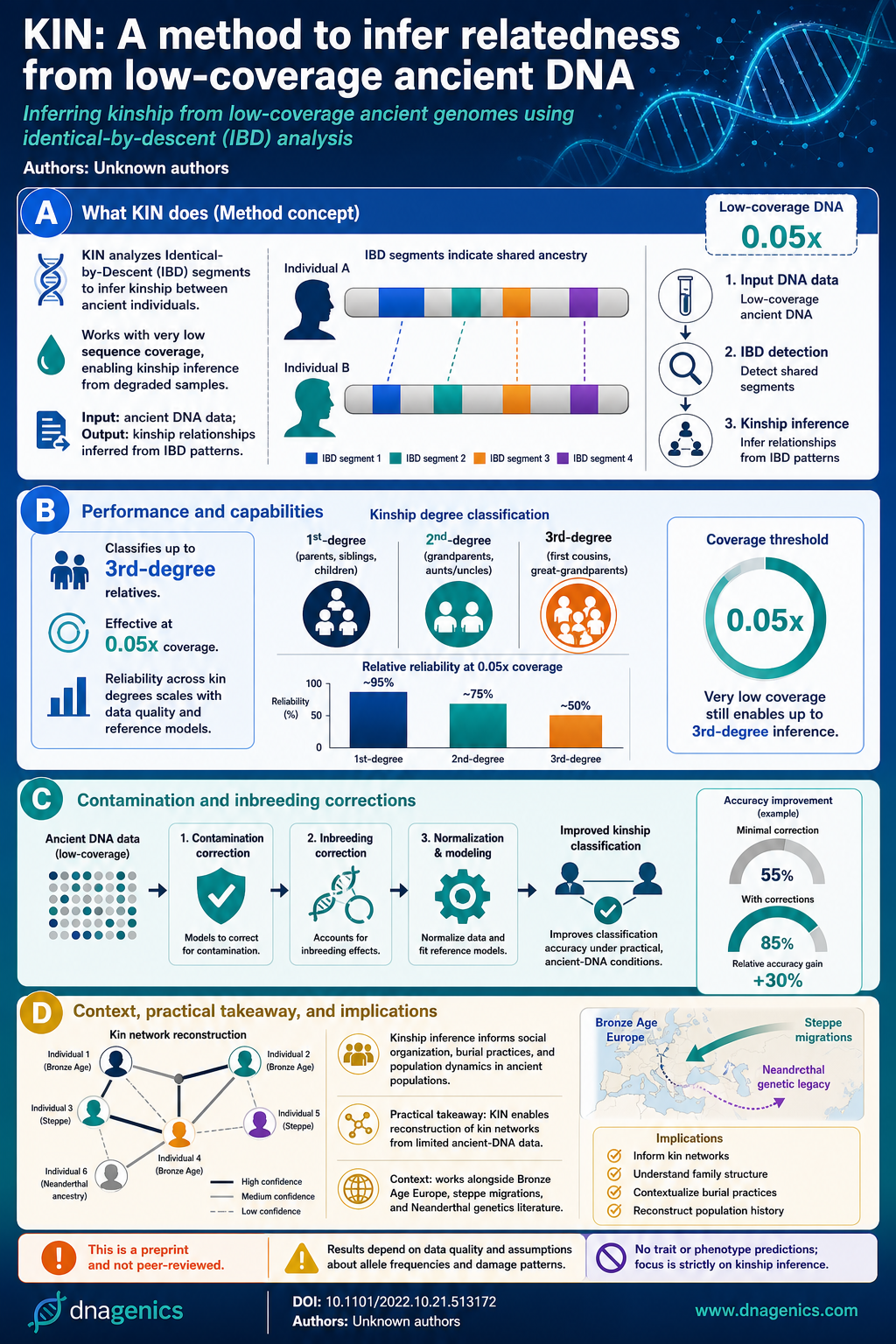

The image helps readers grasp the practical thresholds (e.g., 0.05x coverage) and the decision boundaries between degree classifications, including the distinction between sibling and parent-child relationships. It serves as a quick reference for educators and researchers communicating findings to non-specialists.
Why It Matters
Reconstructing kin networks in ancient contexts sharpens our understanding of how families influenced social organization, burial practices, and population dynamics. By enabling reliable kinship inference from low-coverage data, KIN broadens the range of archaeogenomic datasets that can contribute to questions about ancestry, migration, and cultural exchange. This method also sets a precedent for incorporating contamination and inbreeding-aware models into kinship analyses, which could inform future standards in ancient DNA research and downstream population genetics analyses.
Looking ahead, applying KIN to additional populations and time periods could illuminate how family ties shaped mobility, alliance-building, and demographic shifts. Integrating kinship results with archaeological records and isotopic analyses may yield richer portraits of life in the past and guide new hypotheses about how kin-based social structures formed and dissolved over millennia.
References
- View publication on DnaGenics
- KIN: A method to infer relatedness from low-coverage ancient DNA
- DOI: 10.1101/2022.10.21.513172