Introduction
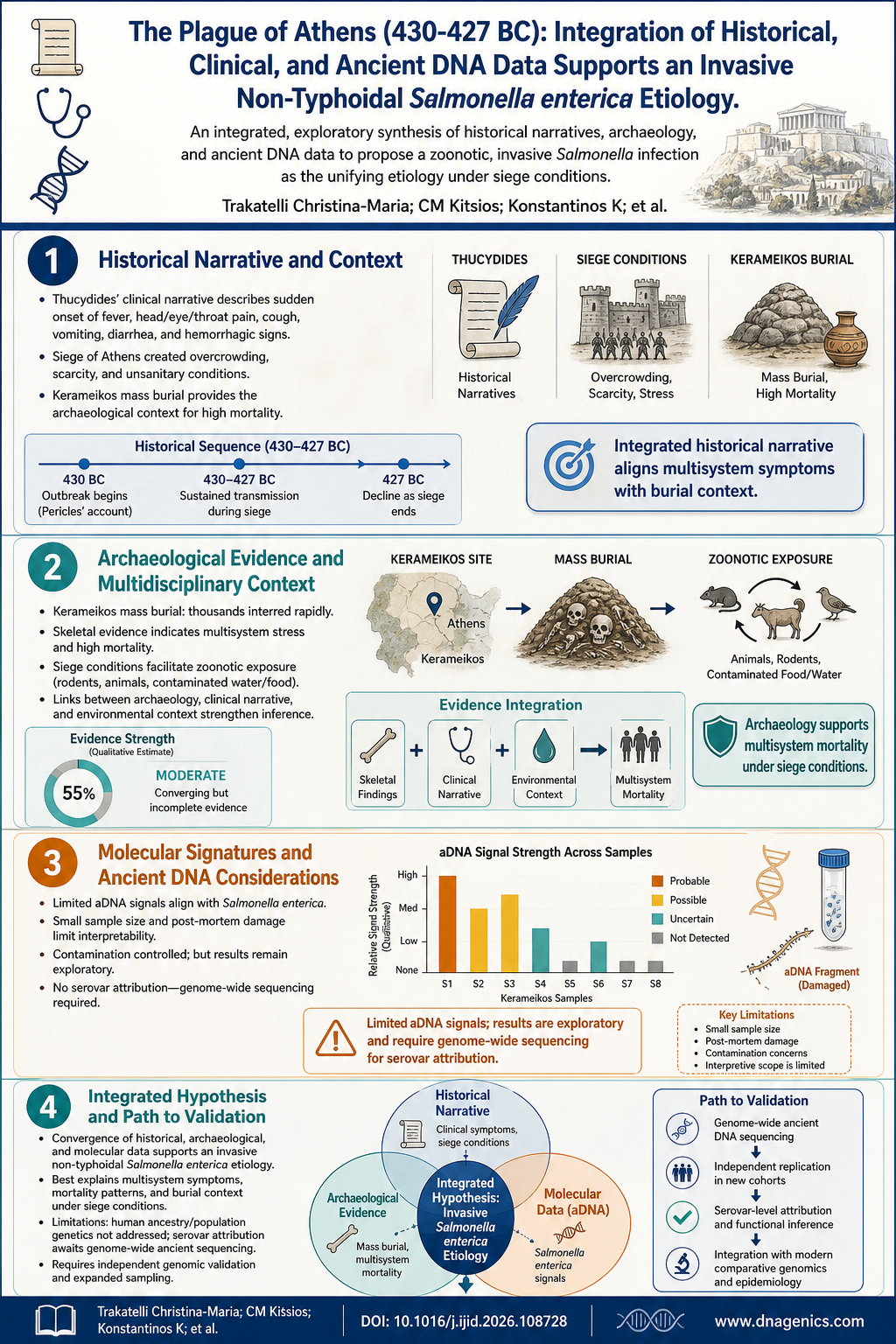
The Ashkenazi Jewish (AJ) population is a well known genetic story in microcosm: a relatively closed community that sits at the crossroads of European and Middle Eastern gene pools. This study sequences 128 high coverage AJ genomes to build a reference panel uniquely tailored to this group, revealing a rich landscape of variants and a history shaped by bottlenecks and expansion. The work matters because a population-specific reference panel dramatically improves variant interpretation in AJ patients and accelerates personalized medicine for individuals with Ashkenazi ancestry.
Why does this research matter beyond cataloging variants? AJ’s genetic distinctiveness provides a powerful lens for disease mapping and population genetics. The panel enhances imputation accuracy for AJ SNP arrays, helps identify variants more likely to influence health in AJ individuals, and offers a clearer picture of how AJ ancestry arose from European and Near Eastern sources. This study also highlights how demographic history and ancient migrations are reflected in modern DNA, informing both ancestry storytelling and clinical genetics.
Key Discoveries
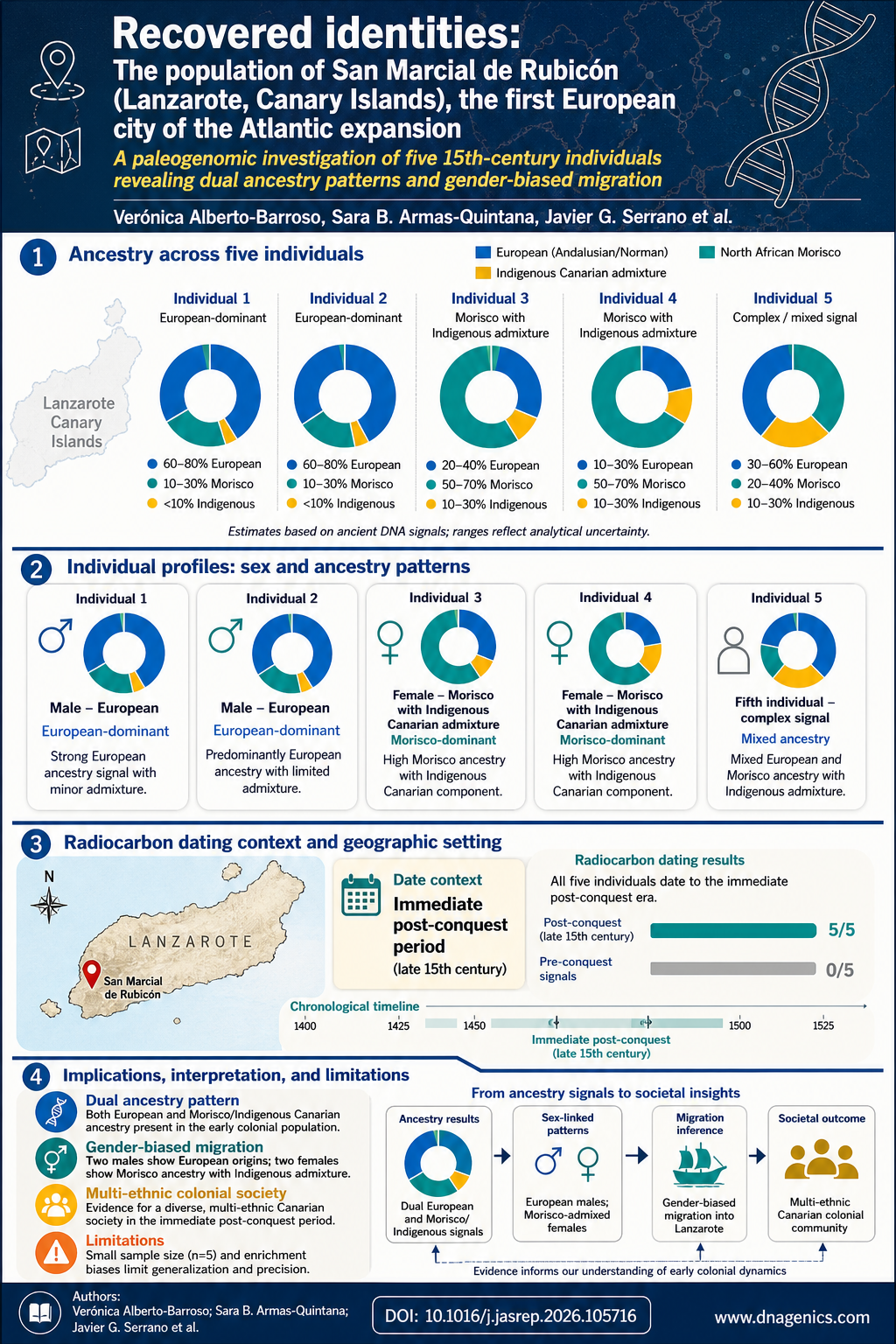
- Ashkenazi Jewish (AJ) population forms a distinct genetic cluster with close links to European and Middle Eastern populations.
- AJ reference panel contains ~47% more novel variants per genome and ~14% more population-specific variants than a European reference panel.
- Imputation accuracy improves by about 28% when using an AJ reference panel, reducing discordance and increasing genotype quality (r^2).
- Long-range IBD sharing within AJ indicates a recent bottleneck and broad haplotype coverage with a small reference panel.
- Admixture history shows AJ as a mixture of European and Middle Eastern origins with ~46–50% European ancestry and divergence times around 12–25 kyr depending on mutation-rate calibration.
What This Means for Your DNA
For individuals curious about their ancestry, the AJ reference panel translates to more accurate genotype imputation for Ashkenazi backgrounds. This means better inference of missing variants in AJ-related studies, improved detection of disease-associated alleles, and more reliable haplotype-based analyses such as ancestry deconvolution and pedigree-based risk assessment. Beginners can think of the panel as a DNA reference more tailored to their lineage, increasing confidence in both ancestry calls and clinical insights, while advanced readers can explore how long IBD tracts reveal shared ancestry and demographic events.
Clinically, the study underscores the value of population-matched references for precision medicine. By enriching for AJ-specific and novel variants, the panel improves filtering of benign variants and highlights disease-associated variants that are particularly informative for AJ patients. Practically, this means clinicians and researchers can interpret AJ genomes more quickly and accurately, supporting personalized screening and therapeutic decisions.
Historical and Archaeological Context
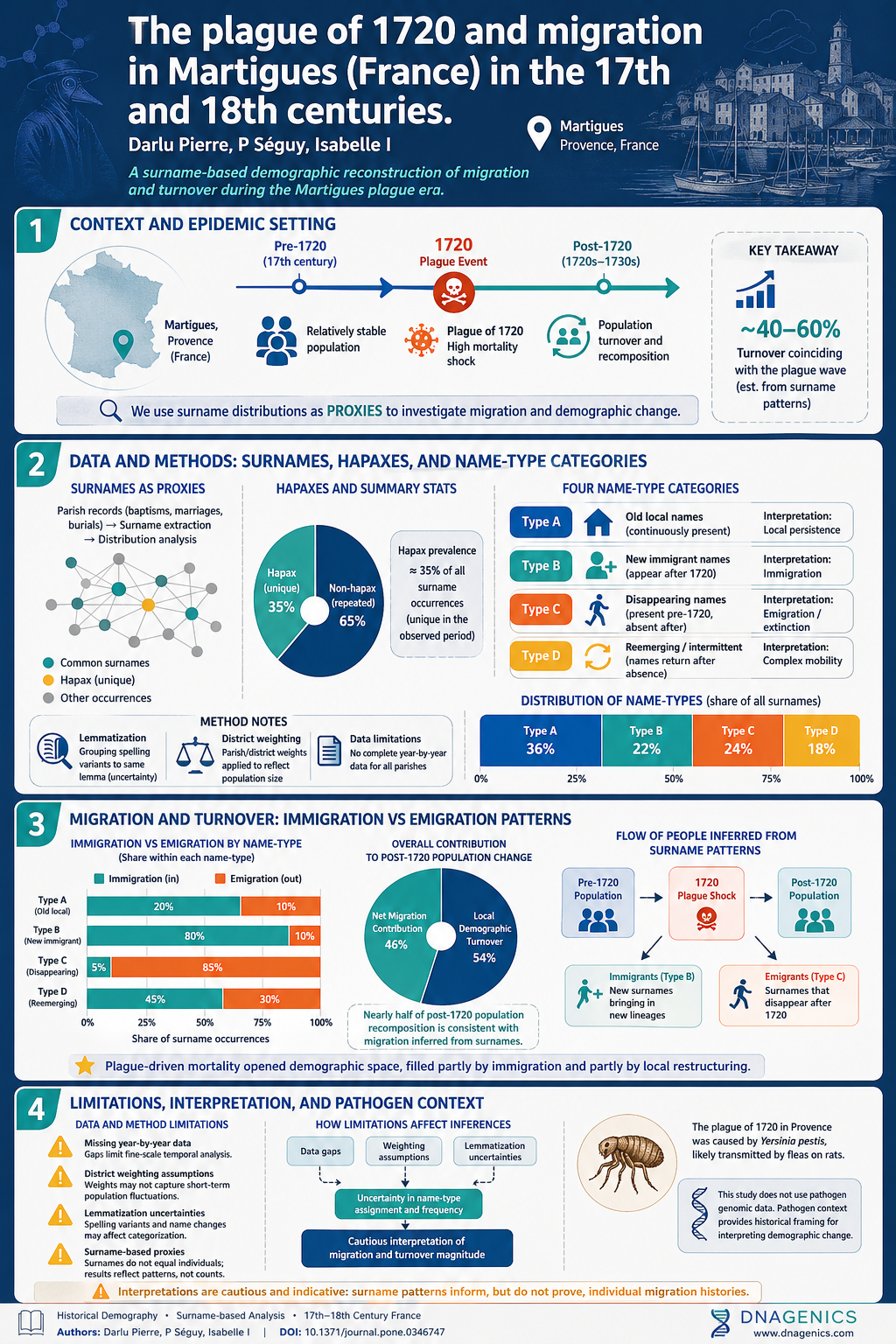
The AJ finding of admixture between European and Middle Eastern lineages fits a complex historical narrative. The joint allele frequency spectrum modeling suggests a post-LGM recolonization of Europe with substantial Near Eastern input, aligning with archaeological and historical records of migrations during the late Pleistocene and early Holocene. The estimated European contribution around 46–50% supports a scenario in which Near Eastern populations contributed to the genetic makeup of European groups, while demographic events such as bottlenecks and rapid expansions shaped present-day variation.
The study also places AJ within a broader timeline of human population dynamics, including deep events dating to 60–90 kyr and a later AJ bottleneck followed by rapid growth. These patterns reflect a sequence of population splits, expansions, and migrations that connect the AJ lineage to both European and Near Eastern contexts, with implications for how cultural and genetic exchange occurred across the region.
The Science Behind the Study
This work rests on high-depth sequencing of 128 complete AJ genomes to build a population-specific reference panel. The authors compare AJ variation against European samples to quantify novel and private variants, providing a benchmark for AJ-focused clinical interpretation. They analyze long identical-by-descent (IBD) segments to infer demographic events and use haplotype-based approaches to leverage the AJ panel for imputation in AJ SNP arrays. The study also models ancient histories for AJ and European populations using joint allele frequency spectra (AFS), yielding estimates of admixture proportions, divergence times, and migration scenarios. Sample size, sequencing depth, and model choice all influence the inferred timelines, illustrating the sensitivity of demographic reconstructions to methodological assumptions.
In Simple Terms: Sequencing 128 Ashkenazi genomes creates a detailed, population specific reference that makes it easier to fill in missing genetic information, interpret disease related variants, and understand how AJ ancestry blends European and Near Eastern roots through shared DNA blocks.
Infographic Section
The study provides an infographic that visualizes AJ population structure, variant richness, and demographic history. It shows AJ as a distinct cluster linked to both European and Middle Eastern groups, highlights long-range IBD sharing, and summarizes admixture proportions and key timepoints.
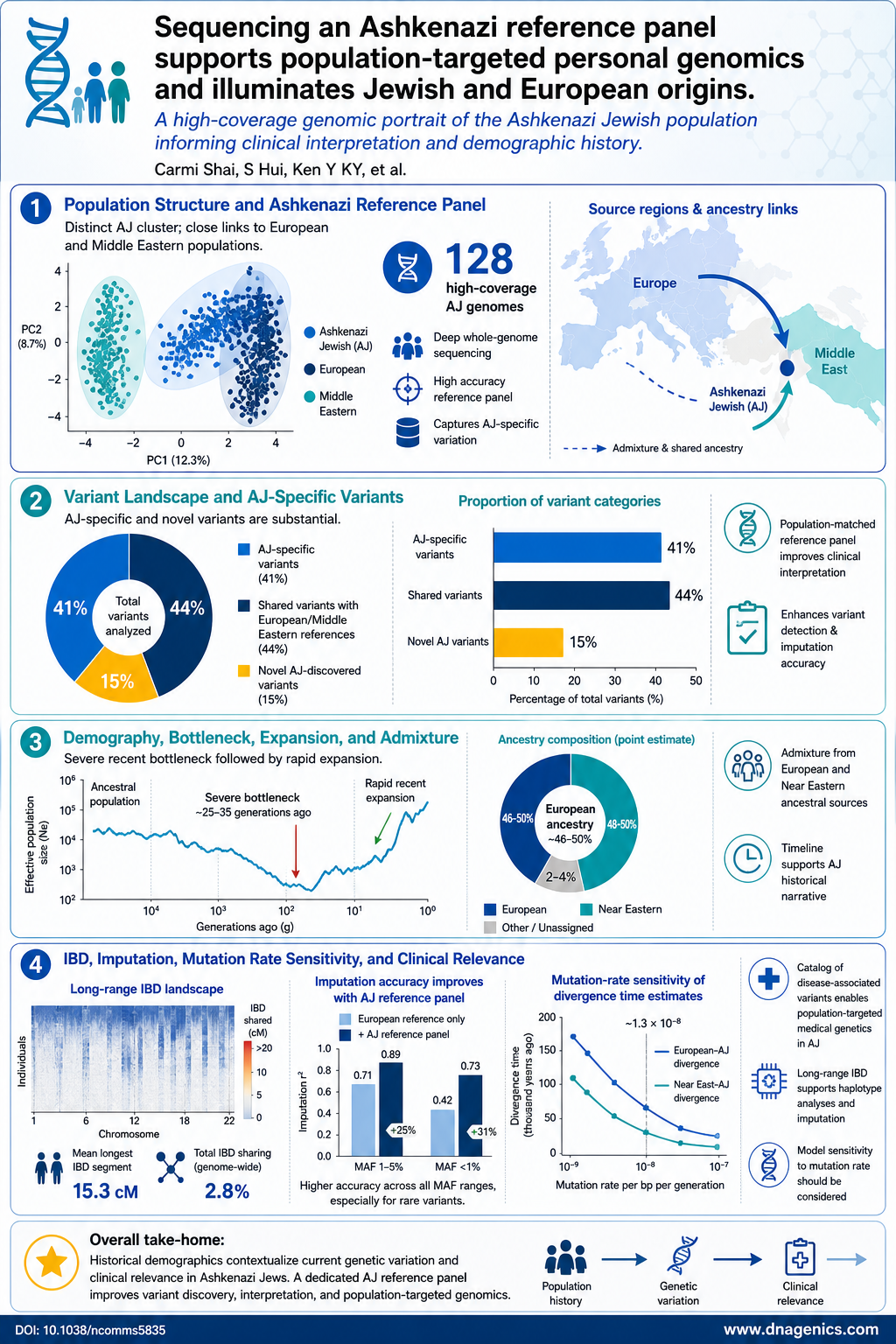
Description: The graphic presents AJ as a unique genetic cluster with connections to European and Middle Eastern populations, highlights extensive IBD sharing within AJ, and depicts estimated admixture proportions and timing that shape present day AJ variation.
Why It Matters
This AJ reference panel advances population genetics by providing a targeted resource for clinical interpretation and genomic medicine. It demonstrates that population matched references can significantly improve imputation, variant filtering, and discovery of disease-relevant alleles in clinically important populations. The findings also emphasize how demographic history, bottlenecks, and admixture shape modern genomes, informing both ancestry narratives and disease risk assessment. Future research could expand to AJ subgroups and other isolated populations, refine mutation rate calibrations, and explore how these panels perform in diverse clinical settings across populations.
References
[Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins] (https://doi.org/10.1038/ncomms5835)