Introduction
The Plague of Athens, described by the ancient historian Thucydides, stands as one of the earliest well documented epidemics in world history. Recent research unites historical narratives, archaeological findings, and ancient DNA data to reassess the cause of this notorious outbreak, which struck Athens during the second year of the Peloponnesian War (430–427 BC). By weaving together different lines of evidence, researchers propose a coherent hypothesis that links clinical descriptions with molecular signals from the dead.
This research matters because it exemplifies how multidisciplinary methods can illuminate ancient disease dynamics and inform our understanding of how epidemics unfold in crowded, stressed populations. The study also demonstrates how ancient DNA (aDNA) analysis can complement traditional historical sources, potentially reshaping debates about the origins and spread of historical plagues. For readers interested in DNA, ancestry, and population history, this work highlights the frontier where pathogens and human history intersect.
Key Discoveries
- Historical movements and siege conditions plausibly shaped epidemic spread and severity under wartime Athens.
- Ancient DNA from Kerameikos indicates Salmonella enterica at the species level; serovar-level resolution remains unresolved due to methodological limits of the data.
- Structured differential diagnosis favors invasive non-typhoidal Salmonella enterica as the most coherent hypothesis among zoonoses when considering zoonotic potential, clinical constellation, and palaeogenomic context.
- Thucydides’ clinical descriptions align with modern iNTS phenotypes (bacteremia, sepsis, encephalopathy) in vulnerable populations, especially under malnutrition and overcrowding.
- Limitations are acknowledged: lack of independent replication, need for genome-wide ancient sequencing, and cautious interpretation of aDNA data.
What This Means for Your DNA
For ancestry buffs, this study showcases how the past can be reconstructed not only from human genomes but also from pathogen DNA preserved in skeletal remains. The integration of clinical narratives with molecular data demonstrates how pathogens leave detectable traces in ancient remains, offering a broader view of population health in historical contexts. While the focus is on a pathogen rather than personal human ancestry, the work underscores the value of archaeogenetics in building holistic pictures of past populations.
In practical terms, readers interested in their own DNA analysis can appreciate how modern techniques are adapted to study ancient diseases. The research highlights that ancient DNA research requires careful interpretation, as signals can be sparse and context dependent. It also reflects why population genetics teams often separate host ancestry from pathogen dynamics, at least in early-stage reconstructions, to avoid conflating human population history with disease ecology.
Historical and Archaeological Context
Thucydides provides a remarkably detailed eyewitness account of the Athens plague, describing multisystem symptoms, rapid spread, and high mortality under siege conditions. The Kerameikos mass burial site offers a crucial archaeological window into this episode, preserving remains that can be tested for ancient pathogens. The convergence of historical narrative and physical evidence situates the outbreak within the broader framework of the Peloponnesian War, urban overcrowding, supply line strain, and sanitation breakdown—the very conditions that can elevate the impact of infectious diseases.
These findings align with known patterns of ancient warfare and city sieges, where sudden population stressors, malnutrition, and close quarters can amplify transmission and disease severity. The timeline (430–427 BC) corresponds to a pivotal phase of the conflict, illustrating how epidemiology and military history intersect in shaping historical outcomes. The geographic focus on Athens and its surroundings provides a contextual roadmap for interpreting similar ancient outbreaks across the eastern Mediterranean.
The Science Behind the Study
The study adopts a multidisciplinary framework, combining historical narrative, archaeology, and ancient DNA analysis. Dental pulp from skeletal remains in the Kerameikos mass burial served as the substrate for pathogen DNA testing, with the goal of identifying species-level evidence for potential etiologies. The results point to Salmonella enterica as the infectious agent at the species level, while acknowledging that serovar-level resolution remains unresolved due to data limitations and the nature of ancient samples. In addition to pathogen signals, the analysis discusses the limitations of aDNA data, including degradation, contamination controls, and the challenges of drawing firm serovar assignments from short or damaged fragments.
The researchers emphasize a structured differential diagnosis that weighs zoonotic candidates against clinical patterns described by Thucydides. They argue that an invasive non-typhoidal Salmonella enterica (iNTS) phenotype best reconciles multisystem symptoms, high mortality under siege conditions, and the involvement of animals in the narrative. Although human ancestry and population genetics are not the focus of this work, the pathogen-centric approach offers a powerful template for how ancient pathogens can be integrated with historical and archaeological data to illuminate past diseases.
In Simple Terms: This study reads like a detective story, using old texts, bones from a mass burial, and tiny DNA fragments to propose a single, testable cause for a famous ancient plague. The evidence points to a dangerous Salmonella infection that spread in crowded, starving cities during war, rather than a purely human-only disease. Researchers stress that more genome-level sequencing is needed to pin down the exact bacterial subtype.
Infographic
The following infographic summarizes the integrated evidence supporting the invasive non-typhoidal Salmonella hypothesis for the Plague of Athens. It connects Thucydides' clinical descriptions with archaeological finds from Kerameikos and the aDNA signals detected in skeletal remains.
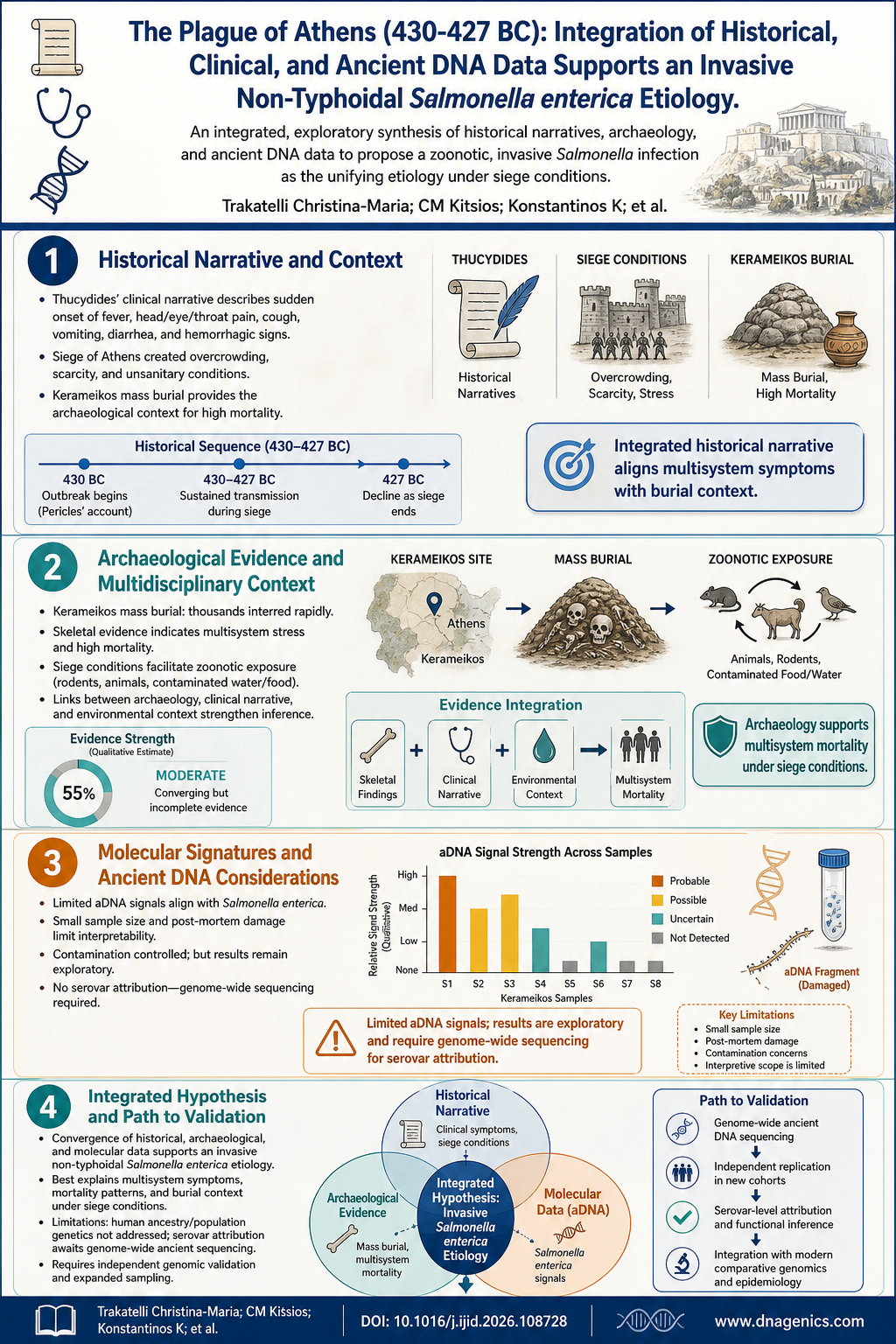

The image illustrates how historical records, burial archaeology, and ancient DNA data converge to form a cohesive hypothesis about the cause of the outbreak, while also highlighting the limitations and areas where future genome-wide sequencing could sharpen the picture.
Why It Matters
This work showcases the power of cross-disciplinary methods to illuminate ancient epidemics and informs how we think about disease dynamics in historical populations. By illustrating a plausible pathogen that coherently explains clinical, archaeological, and molecular data, the study helps set a framework for future archaeogenetic investigations into ancient illnesses. It also highlights the value of transparent acknowledgment of limitations and calls for independent replication and genome-wide sequencing to validate and refine serovar-level attributions.
Looking ahead, advances in ancient genome sequencing and more comprehensive datasets could confirm serovar specifics and expand our understanding of how zoonotic pathogens shaped urban populations in antiquity. The approach demonstrated here may inform future research at the intersection of anthropology, archaeology, and infectious disease genetics, with implications for how we interpret ancient DNA data in the context of human history.