Introduction
Ancient DNA has opened a window into long-lost family ties, social structures, and migrations. Yet decoding kinship from damaged genomes is a delicate business: different methods can yield divergent results depending on how well the data preserve signals of relatedness. The study behind BADGER (Benchmark Ancient DNA GEnetic Relatedness) addresses this by testing six kinship estimation approaches under well-controlled simulations, revealing where reliability breaks down and where conclusions stay sound.
Why this research matters goes beyond method comparison. Kinship inferences influence our understanding of burial practices, social organization, and population movements in archaeology. When samples are few, poorly preserved, or embedded in highly endogamous communities, bias can masquerade as relatedness. This work provides a transparent, reproducible framework to gauge performance, helping researchers interpret ancient DNA findings with appropriate caution and context.
Together, the study situates kinship inference within the broader field of archaeogenetics, where SNP-level data, IBD/IBS signals, and ROH analysis intersect with culture, migration, and demographic history. The BADGER pipeline makes it possible to simulate realistic pedigrees from modern reference panels and generate ancient-like data, enabling robust benchmarking before applying methods to real archaeological contexts.
Key Discoveries
- BADGER provides a rigorous framework to benchmark ancient DNA kinship estimation methods across multiple bias-inducing factors (coverage, damage, contamination, diversity, inbreeding).
- Kinship in ancient datasets is fragile to biases; accuracy cannot be predicted from coverage alone, requiring careful bias mitigation and method selection.
- Historical populations analyzed in related literature (e.g., Late Neolithic mass graves, Bronze Age elites, Danube Basin admixture) illustrate how kinship informs interpretations of ancestry and social structure.
- Genetic markers central to kinship inference include SNP data, IBD/IBS signals, and ROH, with tools like TKGWV2, READv2, KIN, and lcMLkin shaping different approaches to relatedness estimation.
- Cultural context matters: endogamy and inbreeding histories influence relatedness estimates, highlighting the need to integrate archaeological context with genetic results.
- Scientific integrity is upheld through open data resources (BADGER on GitHub/Zenodo, AADR, 1000 Genomes) and explicit discussion of limitations and biases.
What This Means for Your DNA
For people curious about their own ancestry, the study’s takeaways translate into a practical caution: robust kinship inferences from ancient DNA require more than high coverage. Even with modern reference panels, biases from post-mortem damage, contamination, or a population’s history of endogamy can distort relatedness signals. When interpreting ancient kinship results, researchers should consider multiple methods, bias-correction strategies, and archaeological context rather than relying on a single metric.
Practically, this means that a consumer-friendly DNA analysis focusing on ancient kinship should prioritize transparent data processing pipelines, explicit bias assessments, and corroboration with archaeological metadata. The emphasis on multiple bias factors mirrors what advanced ancestry analyses do today: combine genetic signals with historical and cultural context to build a credible family and population narrative.
Historical and Archaeological Context
The benchmarking work connects kinship signals to well-known historical and archaeological scenarios. In Late Neolithic mass graves and Bronze Age contexts, kinship networks can illuminate social organization, inherited status, and ritual practices that shaped the fabric of communities. The study also touches on admixture events in the Danube Basin, where Neolithic farmers and Mesolithic hunter-gatherers interacted, leaving complex genealogical traces that kinship analyses attempt to unravel.
By simulating pedigrees from present-day reference panels (e.g., the 1000 Genomes dataset), the researchers mirror how real ancient populations might carry relatedness patterns across generations. This places methods within a timeline of migration and cultural change, helping to interpret relatedness in light of known demographic events and burial practices.
The Science Behind the Study
BADGER introduces an automated pipeline that first simulates pedigrees using randomly selected present-day individuals from reference datasets, then generates raw ancient DNA sequence data for each member of the simulated family tree. The study then benchmarks several kinship estimation approaches across five biological parameters: sample coverage, post-mortem damage correction, human contamination, genetic diversity, and inbreeding. The goal is to understand how each bias affects accuracy and to provide practical prescriptions for interpreting results.
The methodology emphasizes transparency and reproducibility: simulations rely on publicly available reference panels, and results are anchored in open data resources. The researchers compare multiple widely used kinship tools (including TKGWV2, READv2, KIN, lcMLkin, and BREADR) to assess how different algorithmic assumptions perform under realistic ancient-DNA conditions. Importantly, the study highlights that reliability cannot be inferred from coverage alone, underscoring the need for comprehensive bias assessment in any ancient kinship analysis.
In Simple Terms: This study simulates how families might look across generations using modern DNA, then riffs on how well different methods can recover those relationships when the DNA is degraded like in ancient samples. It shows that data quality and historical context matter as much as raw data quantity when estimating kinship.
[Infographic Section - Infographic available]
The study provides an infographic detailing the BADGER benchmarking workflow, bias factors, and performance outcomes across kinship methods. The image illustrates how pedigrees are simulated, how ancient DNA data are generated under varying conditions, and how method accuracy shifts with coverage, damage masking, contamination, diversity, and inbreeding. It also highlights the key takeaway: reliability depends on multiple interacting biases, not just sequencing depth.
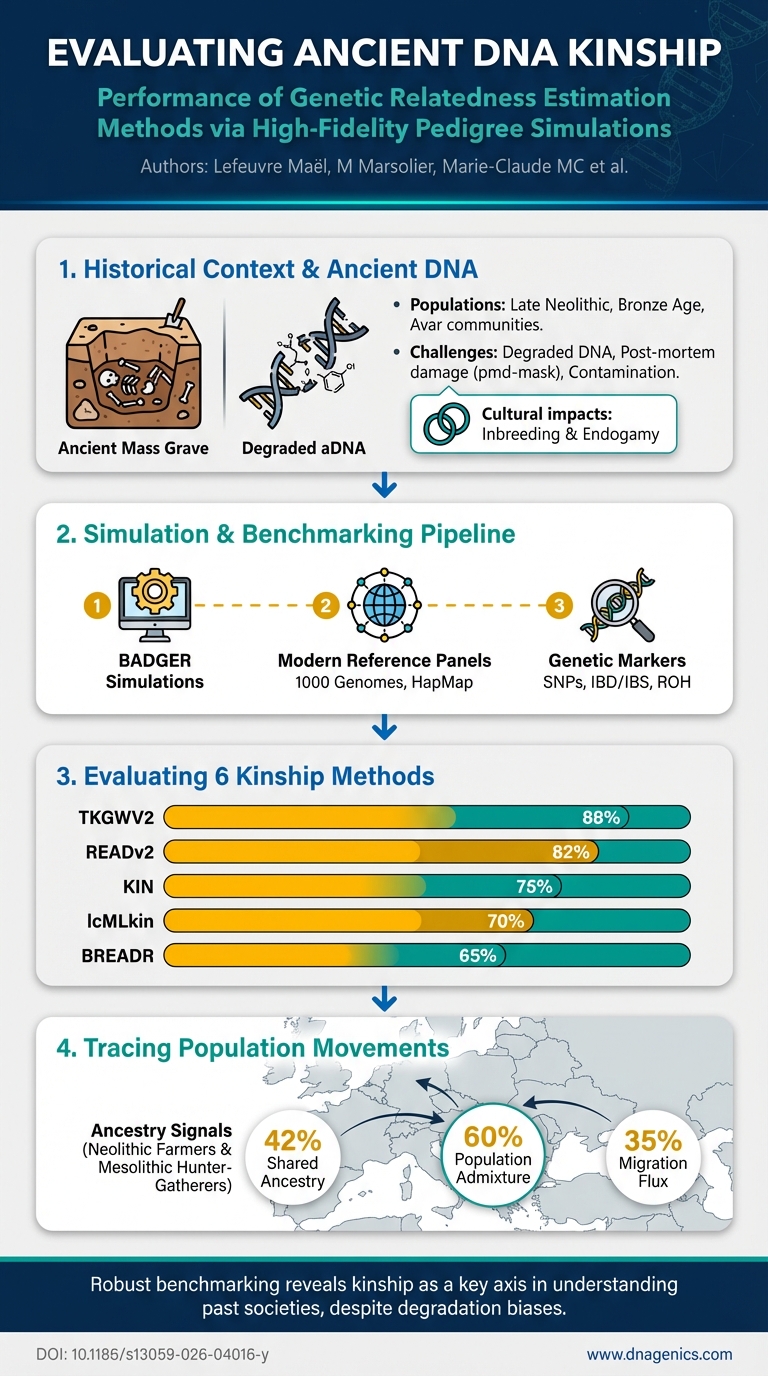
What the infographic shows:
- The multi-factor design used to test kinship methods
- The developmental pipeline from modern pedigrees to ancient-like data
- How performance varies across bias conditions and methods
Why It Matters
This work advances the field by offering a rigorous, transparent framework to benchmark kinship methods in ancient DNA research. It emphasizes that bias-aware interpretation is essential for reconstructing social structure and population history in archaeology. The BADGER approach encourages the community to adopt standardized benchmarking and to integrate archaeological context with genetic results, ultimately strengthening population genetics in ancient contexts and informing future methodological improvements.
Looking ahead, expanding simulations to encompass a broader geographic and temporal range, integrating additional reference panels, and validating findings with real ancient genomes will further enhance our ability to interpret kinship signals in complex past populations.