Introduction
Across the Mediterranean and Sahara, North Africa sits at a genetic crossroads. A new study analyzes 869 complete mitogenomes from six North African populations, with a focus on Morocco, to map maternal lineages and how they moved through time. The work highlights substantial maternal diversity alongside a surprisingly cohesive Maghreb core, suggesting long-standing gene flow within the region and connections beyond Africa.
Why this research matters goes beyond cataloging lineages. Mitochondrial DNA (mtDNA) traces direct maternal ancestry and provides high-resolution windows into historical migrations, population expansions, and region-wide connectivity. By compiling a large, geographically representative set of complete mitogenomes, researchers can resolve deeper time layers than partial sequences allow. Integrating paternal signals from Y-chromosome data (analyzed separately in the study) yields a more complete portrait of North Africa’s population dynamics. This study aligns with broader models of Mediterranean prehistory and historic connectivity while highlighting the limits of uniparental markers for autosomal traits.
The context is clear: Morocco sits at a crossroads of Africa and Europe, and Maghrebi populations have long exchanged genes with neighbors across the Mediterranean. The research draws on uniparental markers to illuminate maternal and paternal lineages, while acknowledging that autosomal data are needed for a fuller, trait-level view of ancestry. The study also reflects modern analytical enhancements, including AI-supported approaches, to parse complex population histories.
Key Discoveries
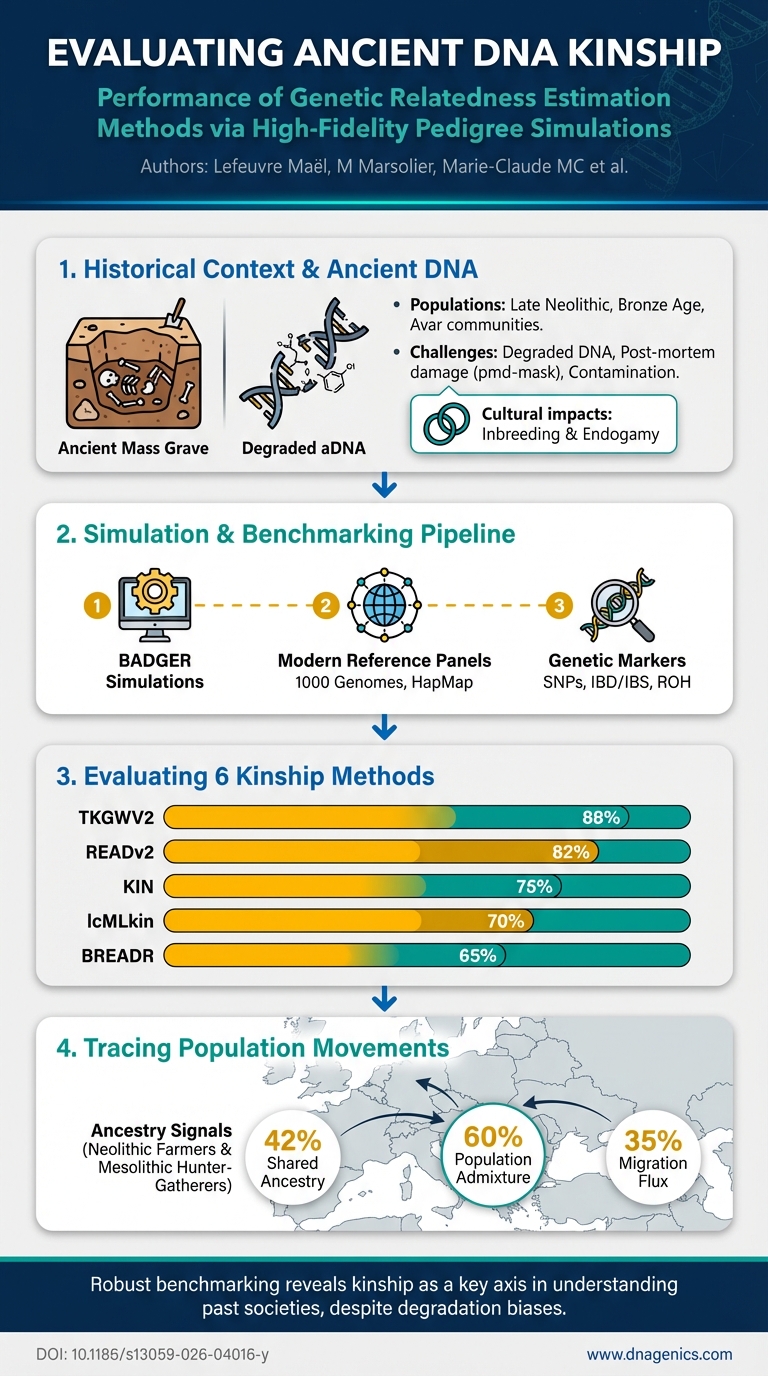
- North African maternal diversity is high yet population differentiation is low (mitogenomes: FST = 0.001–0.014), indicating a cohesive Maghreb core with substantial intra-population variation.
- Dominant maternal haplogroups are U6, H, and L with deep-time diversification in macro-haplogroup L subclades, signaling ancient migrations and back-movements between Africa and Eurasia.
- Morocco shows wide haplogroup distribution across macro-haplogroups, reflecting a history of multiple gene-flow events.
- Paternal North African signal is strong, with E-M81 predominance in Moroccan, Algerian, and Tunisian populations, underscoring a characteristic Maghrebi Y-chromosome background.
- Genetic homogeneity within North Africa is supported by low differentiation measures and AMOVA, reinforcing a cohesive Maghreb core.
- Uniparental markers illuminate ancestry but not phenotype; autosomal genomic data are needed to link to traits and refine time depth.
What This Means for Your DNA
For ancestry enthusiasts, the study suggests that maternal lineages in the Maghreb are richly diverse but regionally cohesive. If your mtDNA falls into haplogroups such as U6, H, or L, you may trace deep-rooted maternal connections to the Maghreb and signals of ancient migrations that shaped North African population history. The Moroccan distributions observed in the study illustrate that multiple gene-flow events—rather than a single donor input—contributed to the present-day maternal landscape.
In practical terms, uniparental markers (mitochondrial DNA and Y-chromosome data) provide powerful anchors for tracing lineage lines and migration stories. However, they reflect only a portion of your ancestry. Autosomal DNA—your overall mix of ancestral segments across the genome—remains essential for a fuller portrait of your ancestry, admixture timing, and trait-related inferences. The study’s maternal perspective, paired with paternal data and autosomal analyses, helps set expectations for what a DNA test can reveal about regional origins and historical connections.
As DNA testing evolves, you can use these insights to interpret your own results more accurately: maternal lineages may reveal deep African and trans-Ma ghreb connections, while paternal signals (like E-M81 in this context) complement that story. Our analytic framework at DnaGenics combines maternal and paternal uniparental data with autosomal information to build a richer, multidimensional ancestry narrative.
Historical and Archaeological Context
The North Africa findings align with a long arc of Mediterranean prehistory and historic connectivity. The presence of deep-time subclades within macro-haplogroup L suggests ancient dispersals from Africa into Eurasia and back-migrations that left lasting imprints on North African mtDNA. This pattern mirrors archaeological and historical records of cross-Mediterranean interactions, including movements linked to climate shifts, trade networks, and cultural exchanges across the Maghreb and Iberian Peninsula.
The analysis reinforces a history of extensive gene flow across the Maghreb, producing a region with high maternal diversity but low between-population differentiation. This genetic cohesion dovetails with archaeological evidence for sustained contact with Europe and the Levant, while maintaining regional continuity within the Maghreb core. The six-population focus—particularly the Moroccan emphasis—helps ground population-genetic models in concrete geography and timeframes, illustrating how demographic expansions and back-migrations shaped modern maternal lineages.
From a broader perspective, the study complements ancient DNA data and archaeological findings by providing a detailed map of maternal lineages across contemporary North African populations. This synthesis strengthens the case for complex, interconnected migration networks rather than simple, one-source ancestry in the Maghreb.
The Science Behind the Study
The researchers compiled 869 complete mitochondrial genomes from six North African populations, with a pronounced focus on Morocco, to examine diversity, haplogroup distributions, and phylogenetic relationships. They used standard population-genetics metrics such as haplotype diversity and neutral tests, finding significant negative values consistent with demographic expansion. Genetic differentiation between populations was low (FST in the range 0.001–0.014), and analysis of molecular variance (AMOVA) indicated that most variation lies within populations, supporting a cohesive Maghreb core.
Phylogenetic and Network analyses (including NeighborNet) revealed that Moroccan sequences are broadly distributed across haplogroups, with deep branching within macro-haplogroup L subclades, indicating ancient migrations and back-migrations. Bayesian demographic analyses showed divergent evolutionary trajectories for prevalent Moroccan haplogroups: expansion of L, decline of U6, and expansion of H. The study also incorporated published Y-chromosome STR data for 2114 North African males, which showed high paternal haplotypic diversity (>0.97) and a strong E-M81 backbone in Moroccan, Algerian, and Tunisian populations. These patterns collectively suggest long-range gene flow and a cohesive Maghreb core, with historic exchanges with Europe and the Levant.
It is important to note that this work relies on uniparental markers, which illuminate ancestry in a direct maternal or paternal line but do not directly translate to autosomal ancestry or phenotypes. To deepen trait, demography, and temporal-resolution insights, autosomal genomic data and ancient DNA are needed. In simple terms, maternal and paternal markers tell distinct, complementary stories about lineage movement, while autosomal DNA provides the broader mosaic of ancestry that shapes who we are today.
In Simple Terms: Maternal DNA follows your mother’s direct line back through time, while the Y-chromosome follows your father’s line. Autosomal DNA is a big, blended record of many ancestors. This study maps maternal and paternal lineages in North Africa, showing a connected Maghreb and multiple migrations, but autosomal data are required to see the full family tree and how traits are inherited.
Infographic Section
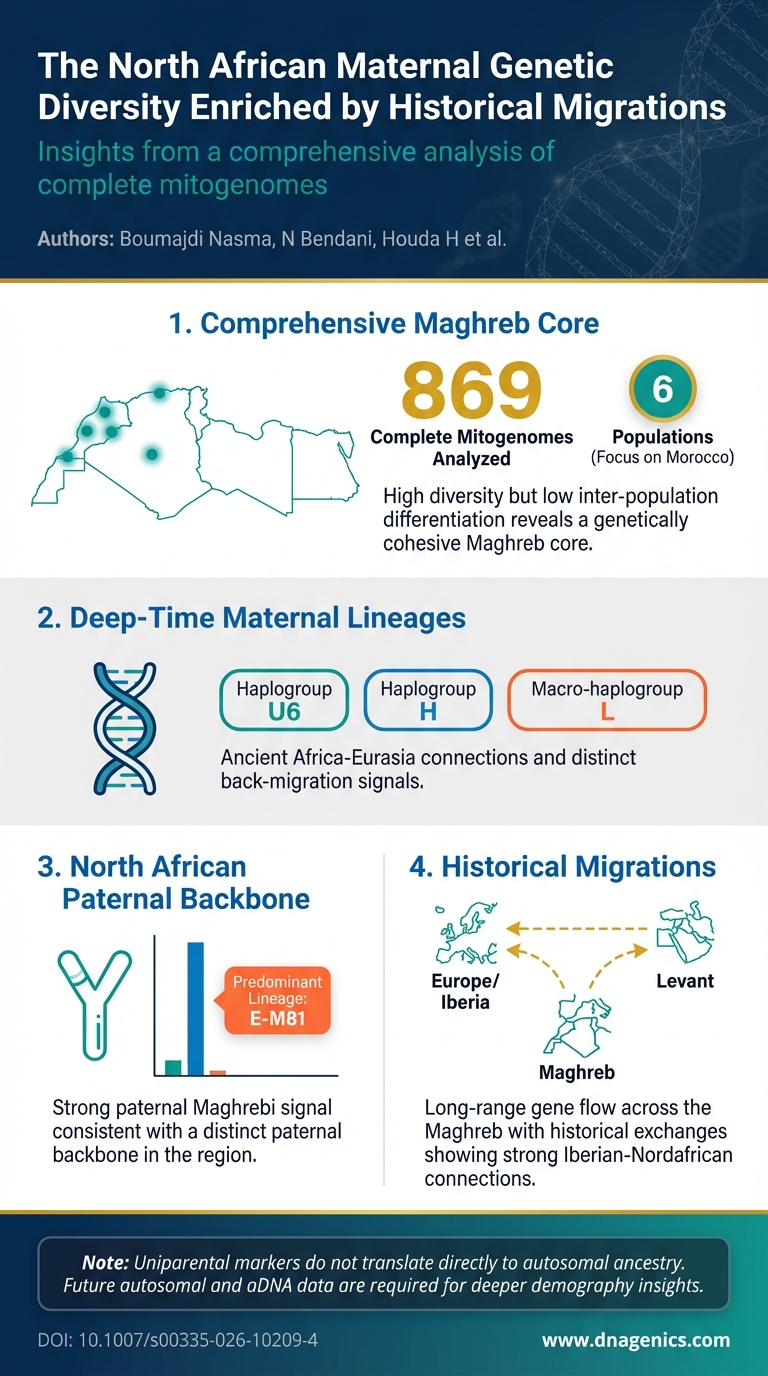
Descriptive text: This infographic summarizes the key maternal-line patterns across North Africa, highlighting the dominant mtDNA haplogroups (U6, H, and L), the low between-population differentiation (FST), and the evidence for a cohesive Maghreb core. It also juxtaposes the strong paternal signal (E-M81) with the maternal patterns to illustrate how maternal and paternal lineages together narrate migration histories in the Maghreb.
Why It Matters
This comprehensive portrait advances our understanding of Maghrebi population history by triangulating maternal and paternal lineages within a cohesive regional framework. The findings reinforce models of ongoing, long-range gene flow across the Maghreb and between North Africa, Europe, and the Levant, while highlighting the historical depth captured by mitochondrial diversity. For researchers, it clarifies the maternal side of North African ancestry and provides a benchmark for integrating uniparental data with autosomal and ancient DNA to refine demographic timelines and migration events. Future work will benefit from expanding autosomal genomic analyses and incorporating ancient North African genomes to sharpen temporal resolution and trait inference.
References
- View publication on DnaGenics
- The North African maternal genetic diversity enriched by historical migrations: insights from a comprehensive analysis of complete mitogenomes
- DOI: 10.1007/s00335-026-10209-4