Introduction
Two pivotal forces shaped early European genomes: the arrival of Anatolian farmers and the longstanding hunter gatherer populations who preceded them. When these groups mixed during the Neolithic transition, their genomes carried the imprints of migration, diet shifts, and changing environments. Local ancestry inference (LAI) is a powerful tool that can map where in the genome each ancestry originates, revealing how selection may have acted on different genetic backgrounds after admixture. This study pushes LAI to the limits by applying six methods to 176 imputed Neolithic genomes from Europe, assessing how well these tools work when reference data are sparse and admixture events are ancient.
Why this matters goes beyond technical benchmarking. Understanding where and how selection acted after admixture helps explain why certain traits spread with the new farming lifestyle, from pigmentation to metabolism. The researchers also highlight how results can vary depending on the method used, especially in complex regions of the genome. This work provides a careful, method aware framework for using LAI with ancient DNA and sets the stage for more robust interpretations of past population dynamics.
Key Discoveries
- Admixture pattern: Neolithic Europeans arose from a two way admixture between Anatolian farmers and Western hunter gatherers, with farmer ancestry generally predominant in early Neolithic genomes.
- Pigmentation adaptation: Robust and replicated signal at
SLC24A5, consistent with lighter pigmentation arising under farmer ancestry in Neolithic Europe. - Dietary adaptation: Strong, replicated signal at
FADS1/2, linked to fatty acid metabolism and dietary shifts after agriculture. - Other candidate loci: Signals at
PER3andIRAK4suggest possible circadian and immune adaptations, but these are less robust and may be confounded by ancestry misclassification. - HLA region caution: Excess hunter gatherer ancestry at the HLA region is not consistently replicated, reinforcing caution about LAI in highly complex regions and emphasizing method sensitivity across the genome.
Additionally, the study notes that X chromosome signals appeared in discovery but did not consistently replicate, likely due to limited power from sparse X data. The authors advocate multi method validation and benchmarking against allele frequency based approaches to strengthen conclusions.
What This Means for Your DNA
For people exploring ancestry today, this work underscores that local ancestry signals reflect deep past admixture and adaptation, but they are highly sensitive to the methods and data used. When LAI points to robust signals at loci like SLC24A5 or FADS1/2, it suggests that ancient populations carrying those backgrounds experienced directional selection after mixing. However, translating these ancient signals to present day phenotypes requires caution, because modern traits result from thousands of generations of evolution and environmental changes.
In practice, if you are analyzing a European ancestry segment, a robust LAI signal at pigmentation or metabolism genes may indicate inherited blocks from farmer ancestry that carried adaptive variants. But be aware that single method results can differ, and replication across methods and datasets strengthens confidence. The study also highlights that regions with complex genetic architecture, like HLA, can yield biased or inconsistent results when using LAI in ancient genomes.
Historical and Archaeological Context
During the European Neolithic transition, large scale movements of people from Anatolia into Europe introduced farming to the continent. This genetic influx mixed with resident Western hunter gatherers, creating a mosaic genome in early farmers. The two way admixture pattern aligns with archaeological records of farming spread from the Near East into Southeastern Europe and beyond, accompanied by dietary shifts from foraging to agriculture and accompanying changes in environment and lifestyle. The timing of admixture, the proportion of farmer ancestry, and the expansion of agricultural societies all leave signatures in the genome that LAI can begin to resolve, though with caveats in data sparsity and reference bias.
The robust signals at SLC24A5 and FADS1/2 fit a narrative where pigmentation and fatty acid metabolism became advantageous in the farming milieu. SLC24A5 is linked to lighter pigmentation, potentially offering advantages under differentUV exposure or social signaling in farming communities. FADS1/2 relates to fatty acid metabolism, reflecting shifts in diet with agricultural reliance on plant and animal fats. The replication of these signals across multiple datasets strengthens their historical interpretation as adaptive responses to Neolithic life in Europe. While additional loci like PER3 and IRAK4 point to possible circadian and immune adaptations, their inconsistent replication urges caution and further validation. The complex HLA region exemplifies how LAI in difficult genomic regions can produce conflicting results, reminding researchers to apply multi-method checks.
The Science Behind the Study
The authors benchmark six local ancestry inference methods using 176 imputed Neolithic genomes, comparing ancestry proportions, tract length distributions, and selection signatures. By explicitly testing six LAI tools in the ancient DNA context, they show that while individual-level ancestry estimates correlate well across methods, tract lengths and inferred admixture times can diverge by more than an order of magnitude. This sensitivity highlights the challenge of dating admixture with LAI when reference panels are small and temporal divergence is large.
To strengthen conclusions, the team replicated analyses across two independent datasets (n = 378 and 1,121) and integrated results across methods. They identify robust ancestry deviations at SLC24A5 and FADS1/2, and note candidate loci PER3 and IRAK4, which require further replication. The findings also stress that LAI results in complex regions like the HLA are particularly susceptible to bias, and thus require careful cross validation with non LAI methods such as allele frequency based approaches (qpAdm, ADMIXTURE) for benchmarking. Overall, the study advances a thoughtful, multi method framework for investigating adaptive admixture in ancient genomes.
In Simple Terms: Local ancestry inference tries to assign each little block of DNA to one ancestral population. In ancient samples, there are fewer reference genomes and more time since the admixture, so guesses can be uncertain. Look for shared signals that show up across several methods and datasets, which makes the conclusion stronger. Block lengths tell us roughly when mixing happened, with longer blocks indicating more recent admixture and shorter blocks pointing to older events.
Infographic Section
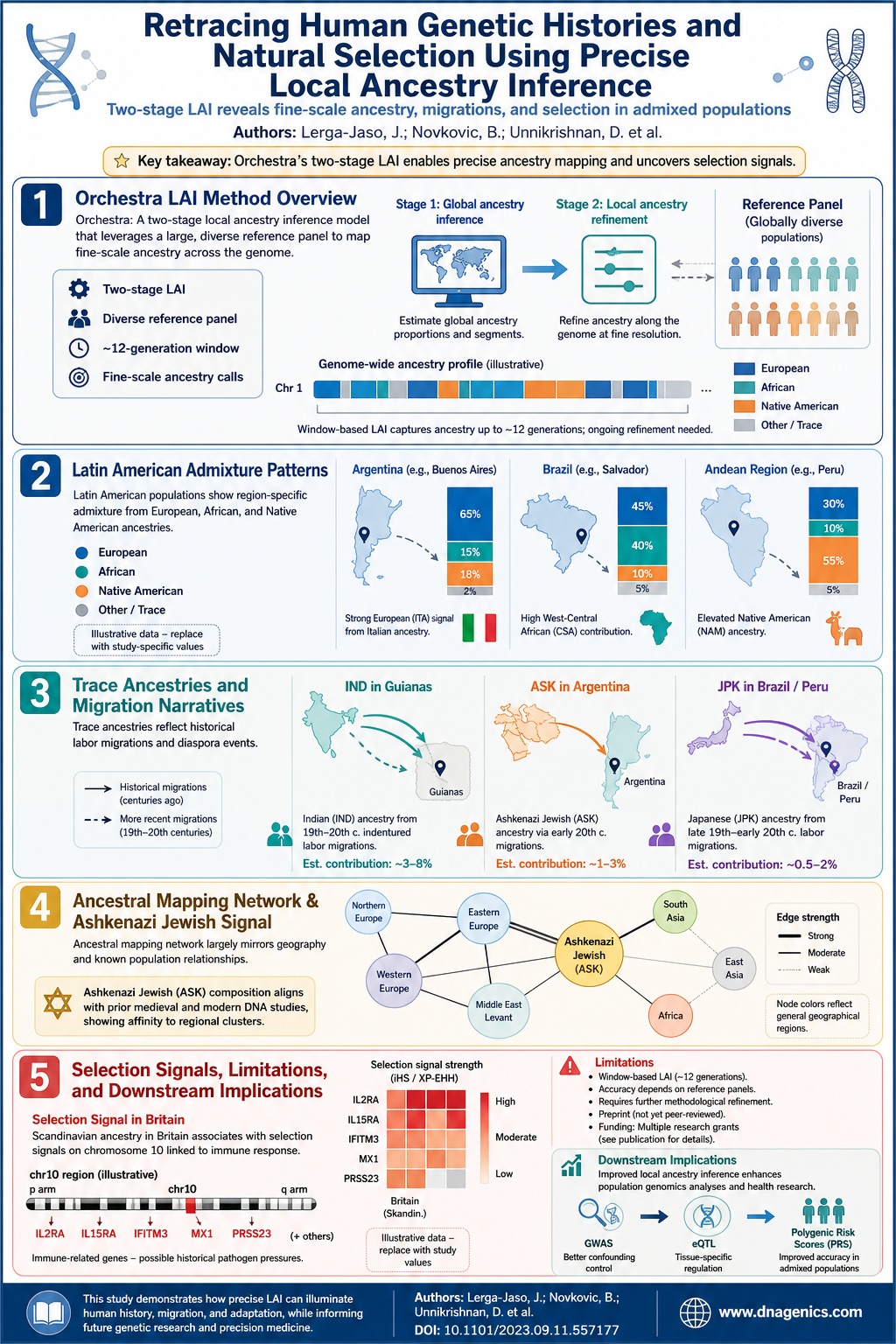
Infographic available: The visual summarizes the LAI workflow, cross method replication, and key findings at SLC24A5 and FADS1/2, plus notes on caution with complex regions like HLA. The image below provides a concise snapshot of how multiple LAI methods converge on robust signals while highlighting areas of discrepancy across methods and datasets.
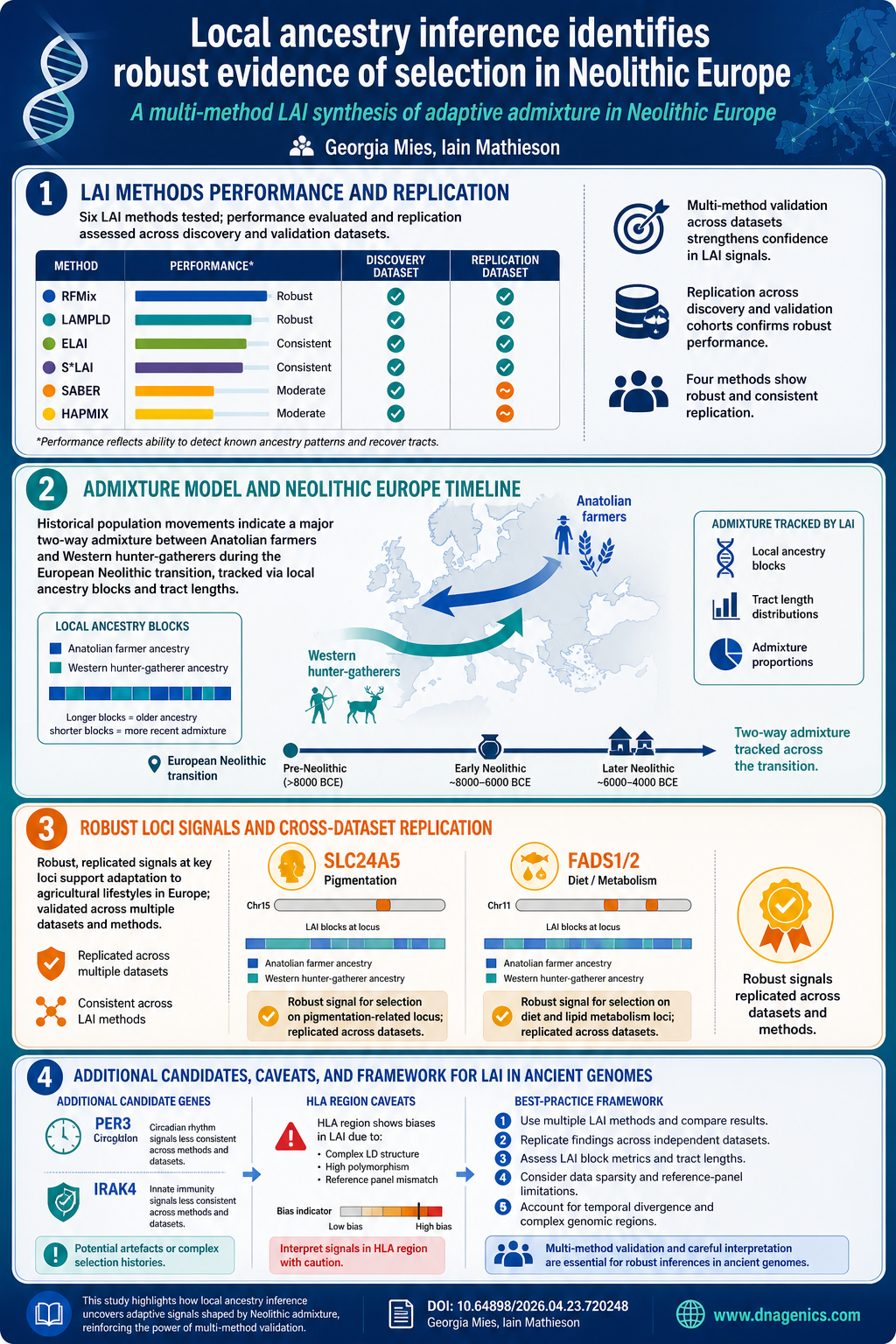

Why It Matters
This work emphasizes that local ancestry inference can uncover meaningful signals of past selection in ancient genomes, but results are highly method dependent. By benchmarking multiple LAI approaches and validating across independent datasets, researchers can gain more reliable insights into how admixture and selection shaped European populations during the Neolithic. The study also points to future directions, including improving reference panels for ancient DNA, refining imputation and phasing strategies, and integrating LAI with allele frequency approaches to build a more robust picture of adaptive admixture. The cautious stance on complex regions like HLA invites ongoing methodological improvements to better handle genomic regions with high polymorphism and complex LD structure.
Future work will likely focus on expanding datasets, enhancing cross method frameworks, and applying these approaches to other ancient contexts to reveal how selection operated after migration, in different environments and times.
References
- View publication on DnaGenics
- Local ancestry inference identifies robust evidence of selection in Neolithic Europe
- DOI